The Chemical Nature of DNA and RNA
The backbone of a nucleic acid is made of alternating pentose ‘sugar’ (deoxyribose sugar in DNA and ribose sugar in RNA) and ‘phosphate’ molecules bonded together in a long chain. Each of the sugar groups in the backbone is attached to ‘nitrogenous base’. The ‘2′-deoxy-’ notation means that there is no -OH group on the 2′ carbon atom
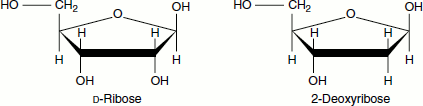
The nitrogenous bases of DNA are adenine, guanine, cytosine and thymine whereas the nitrogenous bases of RNA are adenine, guanine, cytosine and uracil.
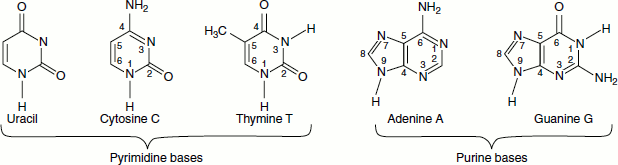
The nitrogenous bases of nucleic are pyrimidines, which are monocyclic, and purines, which are bicyclic. Each has at least one N-H site at which an organic substituent may be attached. They are all polyfunctional bases and exist in tautomeric forms called keto-enol tautomerism. Adenine is 6-amino purine; guanine is 2-amino-6-oxypurine; thymine is 5-methy 1, 2, 4-dioxypyrimidine; cytosine is 4-amino-2-oxypyrimidine and uracil is 2, 4-dioxypyrimidine.
Keto-Enol Tautomerism of the Nitrogenous Bases
Three of the purine and pyrimidine base components of the nucleic acids could exist as hydroxypyrimidine or purine tautomers, having an aromatic heterocyclic ring. Despite the added stabilization of the aromatic ring, these compounds prefer to adopt amide-like structures. The following diagram explains this.
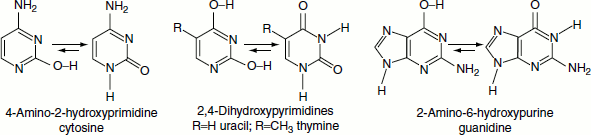
Nucleosides
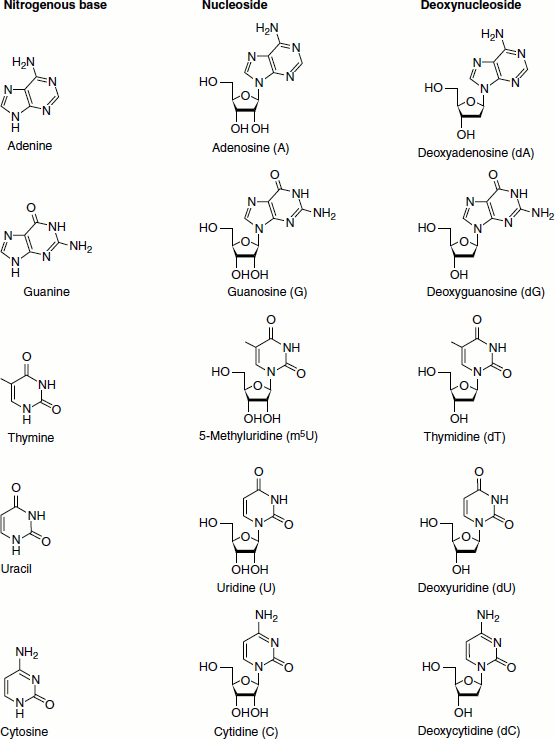
Nucleosides are N-glycosides of 2′-deoxyribose or ribose, combined with the heterocyclic amines through a β-glycosidic linkage. They are formed by the loss of water from a sugar plus a purine or pyrimidine, OH from the anomeric position of the sugar, and H from a nitrogen of the base. Purines bond to the C1′ of the sugar at their N9 atoms. Pyrimidines bond to the sugar C1′ atom at their N1 atoms.
Nucleosides are the basic building blocks of the nucleic acids. In medicine, several nucleoside analogues are used as antiviral or anticancer agents. The nucleosides are adenosine, guanosine, thymidine, methyl uridine, uridine and cytidine respectively named after their bases. The purine nucleosides end with the suffix ‘-sine’: adenosine and guanosine. The pyrimidine nucleosides end with the suffix ‘-dine’: cytidine, uridine and deoxythymidine.
Pyrimidines Exist Entirely in the Anti Conformation
Two conformational variations are possible for nucleosides based on the rotation around the base-to-sugar bond, and puckering of the sugar ring, called ‘Syn’ and ‘Anti’ conformations. Consider the following two structures for adenosine.
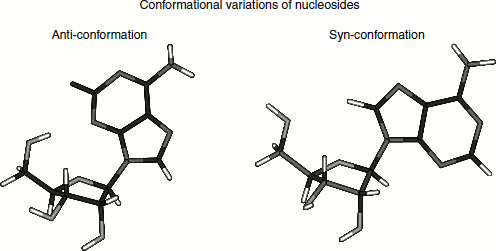
Pyrimidines exist entirely in the Anti-conformation. The puckering of the sugar ring usually involves having either C2′ or C3′ out of the plane formed by C1′, O and C4′.
If C2′ or C3′ is on the same side of the ring as the glycosidic bond, the conformation is described as endo-; if on the other side, it is exo.
Chargaff Rule
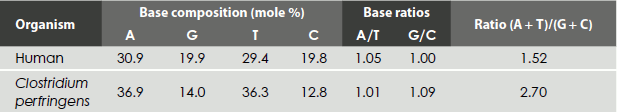
On carefully analysing the DNA from many sources, Erwin Chargaff found its composition to be species-specific. In addition, he found that the amount of adenine (A) always equalled the amount of thymine (T) and the amount of guanine (G) always equalled the amount of cytosine (C), regardless of the DNA source. The ratio of (A+T) to (C+G) varied from 2.70 to 0.35 (Table 1.1).
Table 1.1 Nucleoside base distribution in DNA
Nucleosides are significantly important constituents that are essential for many vital functions. There are two genetic defects related to nucleosides; one is adenosine deaminase (ADA) deficiency and the other is purine nucleoside phosphorylase (PNP) deficiency, account for two immunodeficiencies that result in severe combined immunodeficiency (SCID).
Adenosine deaminase deficiency and purine nucleoside phosphorylase deficiency are autosomal recessive disorders. Adenosine deaminase and purine nucleoside phosphorylase are ubiquitous ‘housekeeping genes’. In both disorders, the enzyme-deficiency results in toxic metabolites accumulation especially in lymphocytes. In adenosine deaminase deficiency, the toxic metabolites block the development of T-cells, B-cells and natural killer (NK)-cells; while in purine nucleoside phosphorylase deficiency, the metabolites are toxic to the development of T-cells.
The inborn errors are characterized by neurodevelopmental delay and are especially prevalent in purine nucleoside phosphorylase deficiency with neurologic symptoms, including mental retardation and muscle spasticity, reported in 67 per cent of patients. Autoimmune disorders, such as autoimmune haemolytic anaemia, immune thrombocytopenia, neutropenia, thyroiditis and lupus also are associated with the disease.
Adenosine deaminase deficiency results in the absence of T-cells, B-cells and NK-cells, resulting in a form of SCID associated with marked lymphopenia. Purine nucleoside phosphorylase deficiency causes decreased numbers of T-cells and lymphopenia. Serum immunoglobulin (Ig) levels are normal to near-normal, but antibodies are deficient.
Nucleotides
Nucleotides are phosphate esters of nucleosides. The phosphoryl group is attached to the oxygen of the 5′-hydroxyl. Monophospates can be further phosphorylated to produce di- and tri-phosphates. The nucleic acid backbone is a polymer with an alternating sugar-phosphate sequence. The deoxyribose/ribose sugars are joined at both the 3′-hydroxyl and 5′-hydroxyl groups to the phosphate groups in ester links, which are also known as ‘phosphodiester’ bonds. Nucleic acids may be formulated as alternating copolymers of phosphoric acid (P) and nucleosides (N), as follows:
∼P–N–P–N′–P–N″–P–N‴–P–N
At physiological pH, the phosphates are ionized, as depicted in the picture.
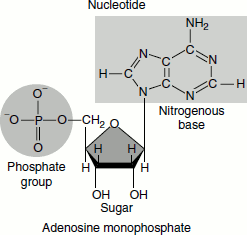
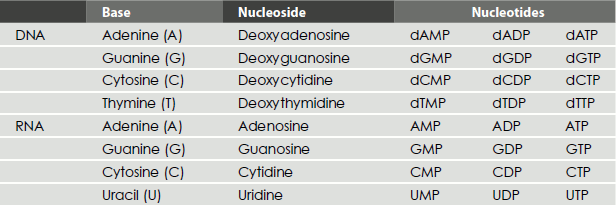
The nucleotides are named by their nucleoside name, followed by the suffix ‘mono-’, ‘di-’ or ‘triphosphate’: for example, adenosine monophosphate, guanosine triphosphate and deoxythymidine monophosphate (Table 1.2).
Table 1.2 Nucleosides and their mono-, di- and triphosphates
Nucleotides have a number of roles. They are
- The monomers for the nucleic acid polymers.
- Nucleoside triphosphates, such as ATP and GTP, are energy carriers in metabolic pathways.
- Nucleotides are also components of some important coenzymes, such as FAD, NAD+ and Coenzyme A.
- Some nucleotides act as intracellular second messengers or signal transducers; for example, cAMP.
‘Single nucleotide polymorphisms’ (SNPs) are DNA sequence variations that occur when a single nucleotide (A, T, C or G) in the genome sequence is altered. For example, an SNP might change the DNA sequence AAGGCTAA to ATGGCTAA.
SNPs do not cause disease, but they can help determine the probability that someone will develop a particular illness. For example, apolipoprotein E (ApoE) is associated with Alzheimer’s disease, which contains two SNPs that result in three possible alleles for this gene: E2, E3 and E4. Each allele differs by one DNA base, and the protein product of each gene differs by one amino acid (genomics.energy.gov, Human genome project).
The Primary Structure of DNA
It is the sequence of nucleotide chains.
Watson and Crick Model of DNA – Secondary Structure of DNA
DNA has a number of special physical and chemical properties that are important to its structure and functioning. The modern era of molecular biology began in 1953 when James D. Watson and Francis H. C. Crick proposed correctly the double-helical structure of DNA (the article ‘Molecular Structure of Nucleic Acids: A Structure for Deoxyribose Nucleic Acid’ was released on 25 April 1953 in the journal Nature), based on the analysis of the x-ray diffraction patterns of DNA fibres taken by Rosalind Franklin and Maurice Wilkins.
The important features of their model of DNA are:
- DNA exists as a pair of molecules and the two strands of DNA are twisted in the shape of a double helix. The two strands are held together by hydrogen bonds, which can be found between the bases attached to the two strands.
- Two helical polynucleotide chains are coiled around a common axis. The chains run in ‘anti parallel direction’. That is, their 5′ → 3′ directions are oppositely oriented.
- The purine and pyrimidine bases are positioned inside the helix, whereas the phosphate and deoxyribose units are on the outside. The planes of the bases are perpendicular to the helix axis. The planes of sugars are nearly at right angles to those of the bases.
- The diameter of the helix is 20 Å. Adjacent bases are separated by 3.4 Å along the axis of the helix and related by a rotation of 36°. Hence, the helical structure repeats after ten residues on each chain; that is, at intervals of 34 Å.
- Adenine always pairs with thymine and guanine always pairs with cytosine.
- Any sequence of bases may occur along a polynucleotide chain. ‘The precise sequence of bases carries the genetic information.’
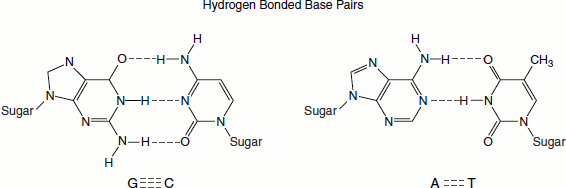
- ‘The base pairing is highly specific. The precise Watson and Crick base pairing of adenine pairing with thymine, and guanine with cytosine, is because of steric and hydrogen-bonding features.’ A is paired with T through two hydrogen bonds and G is paired with C through three hydrogen bonds. This base-pair complementarity is a consequence of the size, shape and chemical composition of the bases. The presence of thousands of such hydrogen bonds in a DNA molecule contributes greatly to the stability of the double helix. Hydrophobic and van der Waals’ interactions between the stacked adjacent base pairs also contribute to the stability of the DNA structure.
- Chemical analysis of DNA (Chargaff, 1950) showed that A equals T and G equals C. In vivo DNA exists predominantly as the B-DNA. Watson and Crick derived model also is based on the B-DNA.
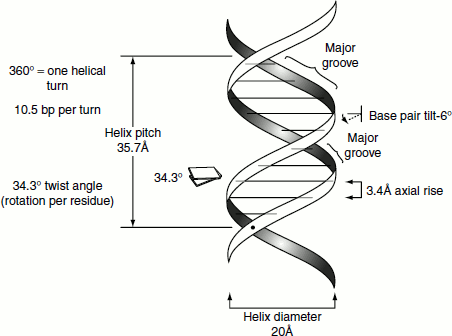
Though many scientific interventions are included to the DNA structure, Watson and Crick model’s four major features remain the same yet today. These features are as follows:
- DNA is a double-stranded helix. The two strands are connected by hydrogen bonds. The A bases are always paired with Ts and the C bases are always paired with Gs. This explains the Chargaff’s rule.
- Most DNA double helices are right-handed; that is, if you were to hold your right hand out, with your thumb pointed up and your fingers curled around your thumb, your thumb would represent the axis of the helix and your fingers would represent the sugar-phosphate backbone. Only one type of DNA, called ‘Z-DNA’, is left-handed. The right-handed helix is the favoured conformation in aqueous systems and has been termed the ‘B-helix’.
- The DNA double helix is anti-parallel, that is one strand runs in the 5′ → 3′ direction, while the other strand runs in the 3′ → 5′ direction. Nucleotides are linked to each other by their phosphate groups, in which the 3′-OH end of one sugar binds to the 5′-PO4 end of the next sugar.
- The outer edges of the nitrogen-containing bases are exposed and available for potential hydrogen bonding as well apart from connecting the two strands. These hydrogen bonds provide easy access to the DNA for other molecules, including the proteins that play vital roles in the replication and expression of DNA.
The DNA Grooves
The twisting of the DNA strands around each other leaves gaps between each set of phosphate backbones. There are two gaps/grooves created because of such twisting around the surface of the double helix: one groove, called the ‘major groove’, is 22 Å wide and the other, called the ‘minor groove’, is 12 Å wide. The edges of the bases are more accessible in the major groove. As a result, DNA-binding proteins such as transcription factors usually make contacts to the sides of the bases exposed in the major groove.
DNA Conformations
The precise geometries and dimensions of the double helix can vary. DNA can exist in three different conformations namely A-, B- and Z-DNAs.
B-DNA
The most common conformation in most living cells (Wat-son and Crick Model) is known as B-DNA (Figure 1.1).
A-DNA
A-DNA is a shorter and wider form. It has been found in dehydrated samples of DNA and rarely under normal physiological circumstances. The A-form is more compact than the B-form. There are 11 bases per turn and the stacked bases are tilted (Figure 1.1).
Figure 1.1 Different forms of DNA
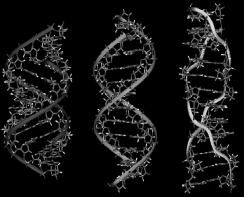
Z-DNA
The Z-DNA helix is left-handed and has a structure that repeats every two base pairs. Unlike A- and B-DNAs, there is a little difference in the width of the major and minor grooves. The formation of this Z-DNA conformation is generally unfavourable. However, certain conditions can promote it. Alternating purine–pyrimidine sequence (especially poly(dGC)2) or high salt and some cations (all at physiological temperature, 37°C, and pH 7.3–7.4) are some of the factors that favour Z-DNA conformation (Table 1.3).
Z-DNA is a transient form of DNA and it can exist only occasionally, in response to a certain types of biological activity. Z-DNA was first discovered in 1979, but its existence was largely ignored until recently. Certain proteins bind very strongly to Z-DNA, suggesting that Z-DNA plays an important biological role in protection against viral disease (Figure 1.1).
Triple Stranded DNA
A triple-stranded DNA structure can also exist in vitro and possibly during the recombination and DNA repair. For example, when synthetic polymers of poly(A) and polydeoxy(U) are mixed, a triple-stranded structure is formed. The synthetic oligonucleotide can insert as a third strand and binds in a sequence-specific manner called ‘Hoogsteen base pairs’ (Figure 1.2).
Cruciforms or Holliday Junction
‘Palindromes’ are words, phrases or sentences that are the same when read forward or backward, such as ‘radar’, ‘Madam, I’m Adam’ etc. DNA sequences that are ‘inverted repeats’, or palindromes, can form a tertiary structure known as a ‘cruciform’ (meaning ‘cross-shaped’) if the normal inter-strand base pairing is replaced by intra-strand pairing. In effect, each DNA strand folds back on itself in a hairpin structure to align the palindrome in the specific base-pairing form resulting in the formation of cruciforms. The unpaired DNA sequences are looped as the result they are never as stable as normal DNA duplexes. Cruciform structures have a twofold rotational symmetry about their centres and these regions act as the recognition sites for specific DNA-binding proteins. This structure is important for the critical biological processes of DNA recombination and repair that occur in the cell (Figure 1.3).
Table 1.3 Comparison of properties of different forms of DNA
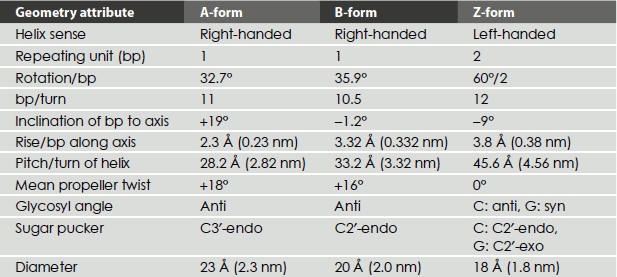
Figure 1.2 Triple stranded DNA
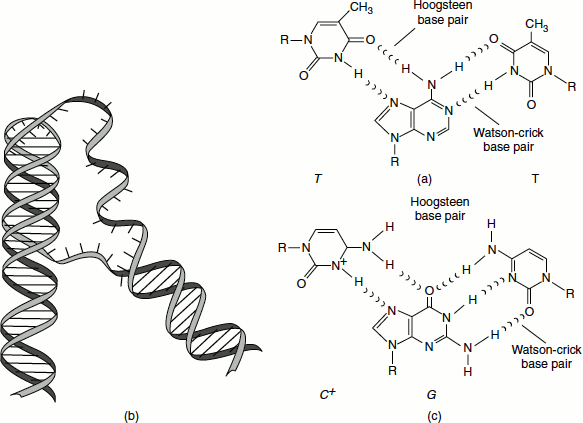
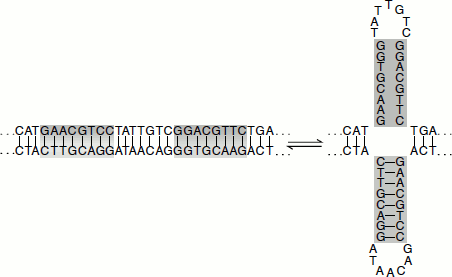
Figure 1.3 Cruciforms or holliday junction
The formation of a cruciform structure from a palindromic sequence within DNA. The selfcomplementary inverted repeats can rearrange to form hydrogen-bonded cruciform loops.
Tertiary Structure
This refers to how a DNA is stored in a confined space to form the chromosomes. This varies in prokaryotes and eukaryotes. In prokaryotes, the DNA is folded like a super-helix, usually in circular shape and associated with a small amount of protein. The DNA of cellular organelles such as mitochondria and chloroplasts also takes a similar structure. In eukaryotes, since the amount of DNA from each chromosome is very large, it is compacted into the nucleus with the help of proteins such as histones and other proteins of non-histone nature.