The double-stranded structure of a DNA determines its biological function. Metabolic events involving unwinding impose great stress on the DNA because of the constraints inherent in the double helix. Molecular processes such as replication and transcription require the unwinding of the DNA double helix. While unwinding, correct topological tension in the DNA (super-helical density) should be maintained in order for genes to be regulated and expressed normally.
In 1965, Vinograd et al. discovered that the circular DNA chromosomes isolated from small viruses such as SV40 or polyoma virus were in a highly compact or folded conformation.
This supercoiling or writhing of the circular DNAs was a result of the DNAs being under wound with respect to the relaxed form of DNA.
When a linear DNA is free in solution, it assumes a pitch that contains 10.4 base pairs per turn. As the result, the DNA is less tightly wound than 10.0 base pairs per turn in the Watson and Crick B-form DNA.
In order to understand the origin of supercoiling; imagine a linear DNA of 4,200 base pairs in length. If the DNA were in the B-form, one would expect the two strands of the helix to be wrapped around each other 400 times (4,200 bp/10.4 bp/turn). Imagine a linear DNA in which the two ends become connected to form an open circle. This is referred to as a relaxed circular DNA. On the other hand, if the linear DNA were unwound 10 per cent, say 40 turns, before its ends were joined, then the DNA molecule would be under stress. When the molecule is free in solution, it will coil about itself in space, as the two strands simultaneously twist about each other in order to return to equilibrium value of 10.4 base pairs per turn.
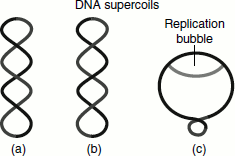
The DNA that is ‘underwound is referred to as negatively supercoiled’. The DNA in this case forms the right-handed double helices. The DNA that is ‘overwound is referred to as a positively supercoiled DNA helix and is a left-handed helix’ (Figure 1.5).
Figure 1.5 The structure of super-coils. (a) Positive supercoils – the front segment of a DNA molecule cross over the back segment from left to right (b) Negative supercoils (c) The positive supercoil in bacteria during DNA replication
Linking, Twisting and Writhing
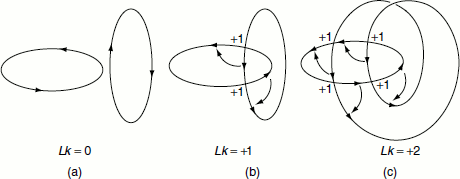
The total number of times one strand of the DNA helix is linked with the other in a covalently closed circular molecule is known as the ‘linking number (Lk)’ (Figure 1.6).
Figure 1.6 Linking, twisting and writhing
Salient features about linking number
- The Lk is only defined for a covalently closed DNA and its value is fixed provided the molecule remains covalently closed. The Lk does not change whether the covalently closed circle is forced to lie in a plane in a stressed conformation or it is allowed to supercoil about itself freely in space.
- The Lk of a circular DNA can only be changed by breaking a phosphodiester bond in one of the two strands, allowing the intact strand to pass through the broken strand and then rejoining the broken strand.
- Lk is always an integer since two strands must always be wound about each other an integral number of times upon closure.
- The Lk of a covalently closed circular DNA can be resolved into two components called ‘the twists (Tws) and the writhes (Wrs)’. Lk = Tw + Wr
- In a relaxed circular DNA duplex of 500 bp, L is 50 (assuming that 10 bp per turn in the B-DNA).
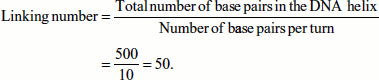
- The Lk for a relaxed DNA is usually taken as the reference parameter and is written as L0.
- By convention, the Lk is defined as positive for right-handed helix and negative for left-handed helix. Since the left-handed Z-DNA occurs very rarely negative Lks are not encountered in all DNA studies for practical purpose.
Twists and writhes
‘The number of times the two strands of DNA are twisted about each other is called the twists (Tws). The writhes (Wrs) are the number of times that the DNA helix is coiled about itself in three-dimensional space.’ Tw and Wr are geometric rather than topological properties. The Tw and the Wr are not necessarily integers. It is just their sum, the Lk that is an integer.
If we use an SV40 DNA molecule, for example, which is precisely 5,243 base pairs in length, we would find that:
Lk = Tw + Wr
Lk = 5,243/10.4 = 504.13.
The DNA length and its pitch in solution determine the Tw of DNA.
Tw = length (bp)/pitch (bp/turn).
The Tw and the Lk determine the value of the Wr.
Wr = Lk − Tw
= 504.13 − 24.13
= 480.
Unlike the Tw and the Lk, the Wr of a DNA only depends on the path the helix axis takes in space. If the path of the DNA is in a plane, the Wr is always zero. In addition, if the path of the DNA helix were on the surface of a sphere, then the total Wr can also be shown to be zero.
Wrs can come in different forms. If a DNA molecule wrap around itself, then the Wrs are known as supertwists. If a DNA molecule wrap around something else (another molecule for instance), then the Wrs are known as ‘solenoidial’ Wrs. In solution, the Wrs can isomerize between the supertwist and solenoidal forms.
Measuring supercoils
The topoisomerases change the Lk (some directly and some indirectly). The change in the Lk, ΔLk, is a measure of the supercoiling. If the Lk in a supercoiled DNA and the Lk in the relaxed state (both of which must be integers) is compared, then the ratio would be:
ΔLk/Lk = σ = the superhelical density.
‘Super helix density or specific linking difference’ is the difference between the Lk of a DNA in the supercoiled form and the Lk in the relaxed form.
A σ of 0.1 means that 10 per cent of the helical turns in a sample of DNA (in its B configuration) have been removed. This underwinding results in negative supercoiling. In a cell, σ is usually of 5–7 per cent.
The superhelical density of a circular DNA can be observed and measured in several ways; for example, electron microscopy, sedimentation velocity or electrophoresis. Supercoiling can be measured by sedimentation procedure. Since supercoiled molecules are more compact, they sediment faster in a centrifuge than when they are relaxed. Supercoiling can also be determined by electrophoresis in an agarose gel. A supercoiled DNA migrates much more rapidly than does a relaxed molecule of the same length. The DNA separates into discrete bands depending on the Lk. Since the DNAs resolved in this way differ from each other only in their topology, they are referred to as ‘topological isomers or topoisomers’. Molecules that differ by one unit in Lk can be separated by electrophoresis in agarose due to the difference in their Wr (that is due to difference in folding).
Topoisomerases
‘Topoisomerases’ are enzymes that change the Lk of a circularly wound double-stranded DNA. The change in Lk changes the Wr. The variation in Wr subsequently changes the state of the compaction of the DNA molecule. The naturally occurring DNA is underwound or negatively supercoiled. This is advantageous because it permits the DNA to be transiently and locally melted to permit the enzymes of the DNA replication and transcription to copy and synthesize new DNA or RNA.
There are two classes of topoisomerase: Type I and Type II.
Type I topoisomerases
These enzymes remove supercoils by breaking only one of the two strands of the DNA. As a result, these enzymes change the Lk by 1 each time. The best-characterized member of this class in E. coli is Topoisomerase I. This enzyme is 864 amino acids in length and is monomeric; it is encoded by the topA gene.
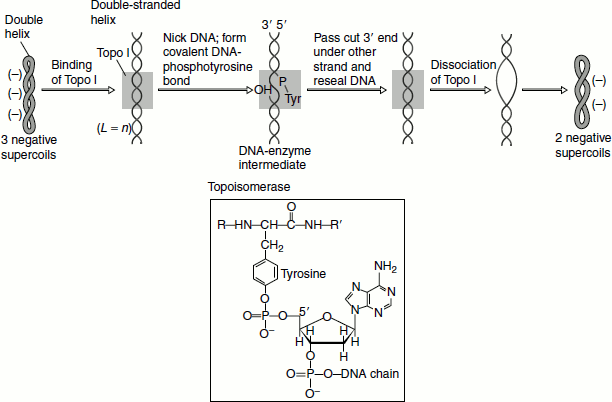
The mechanism of catalysis involves the formation of a covalent intermediate between a tyrosine residue and the phosphodiester backbone. Specifically, nucleophilic attack from the hydroxyl group of tyrosine to a phosphorus atom creates a phosphodiester link between the enzyme and the DNA and generates a free 5′-hydroxyl group. Formation of this bond is energetically neutral as the reaction involves the replacement of one phosphomonoester bond with another—such reactions are called ‘transesterification’ reactions. The other strand of the DNA is held in place by binding non-covalently to a domain of the enzyme. The cleaved strand is resealed. Thus, one supercoil is removed. Topo I from E. coli acts only on negative supercoils; while eukaryotic Topo I can remove both negative and positive supercoils (Figure 1.7).
Figure 1.7 Topoisomerase I
Type II topoisomerases
These enzymes act through a mechanism, in which both the phosphodiester backbone chains are broken simultaneously. As a result, the Lk changes by two. Some Type 2 enzymes can use ATP to introduce the superhelical turns into the DNA. The best-characterized member of this class is E. coli Topoisomerase II—better known as DNA gyrase. E. coli DNA gyrase is a tetrameric protein consisting of two A subunits (875 aas) and two B subunits (804 aas). Depending on the DNA substrates, these enzymes can change positive supercoils into negative supercoils or increase the number of negative supercoils by 2. Type II topoisomerases catalyse catenation and decatenation, i.e., linking and unlinking of two different DNA duplexes. The enzyme also introduces negative supercoils at or near the Ori C site in the DNA template. DNA gyrase also removes the positive supercoils that are formed ahead of the growing fork during replication (discussed in Chapter 3).
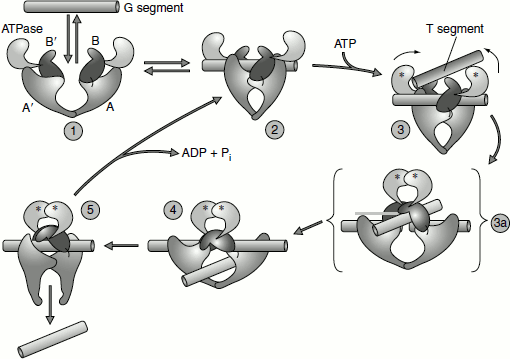
DNA gyrase is composed of two identical subunits. The hydrolysis of ATP by gyrase’s inherent ATPase activity powers the conformational changes that are critical to the enzyme’s operation. The enzyme is a dimer, which has two identical subunits. Initially, the enzyme binds one part of a DNA strand, the G segment, inducing a conformational change in the B, B′, A and A′ domains of the enzyme 2. After the binding of ATP (indicated by the asterisks) and another part of the DNA strand, the T segment, a series of reactions occur in which the G segment is cut by the A and A′ domains of the enzyme and the ends of the G-DNA become covalently linked to tyrosine residues in these domains 3 and 3a. Simultaneously, the ATP-binding domains move towards each other, transporting the T segment through the break and into the central hole 4. The cut G segment is then resealed and the T segment is released by a conformational change that separates the A and A′ domains at the bottom of the enzyme 5. The interface between the A and A′ domains then re-forms, a reaction that requires ATP hydrolysis and regenerates the starting state 2. At this point, the G segment can dissociate from the enzyme by the conversion of 2 into 1. Alternatively, the enzyme can proceed through another cycle, again passing the T segment through the G segment and thus removing two more supercoils (Berger et al., 1996) (Figure 1.8).
Figure 1.8 Topoisomerase II
Topoisomerases are essential enzymes. The mutations of any of the genes coding for topoisom-erases are usually lethal. They are, therefore, the targets for the antibiotics and other drugs. Bacteria can be killed by novobiocin or nalidix acid. Both of these inhibit DNA gyrase. Novobiocin blocks ATP binding and nalidixic acid blocks the breakage and rejoining mechanism. These antibiotics do not inhibit eukaryotic topoisomerases and can be used to eradicate bacterial infections. Some bacteria, however, are now resistant to novobiocin. Eukaryotic topoisomerase inhibitors, such as doxorubicin and etoposide, are used as chemotherapeutic agents.