Prokaryotic DNA Polymerases
DNA polymerases are the enzymes responsible for the synthesis of DNA during replication.
There are three types of prokaryotic DNA polymerases; namely, DNA polymerase I (DNA Pol-I), DNA polymerase II (DNA Pol-II) and DNA polymerase III (DNA Pol-III).
DNA Pol-I
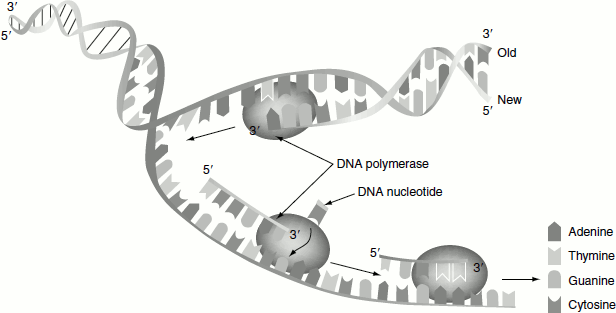
It is the first known DNA polymerase, which was discovered by Arthur Kornberg in 1956 and hence also called Kornberg enzyme. It is a monomeric 928-residue polypeptide. It couples dNTPs on DNA template in a reaction that occurs through the nucleophilic attack of the growing DNA chains 3′-OH on the α-phosphoryl group of the incoming nucleotide. The reaction is driven by the elimination of the pyrophosphate and its subsequent hydrolysis. The error rate of polymerase I in copying the template is very low about one wrong base pair per 10 million. Polymerase I is a ‘processive’ enzyme, i.e., it catalyses a series of successive polymerization steps without releasing from the template. Polymerase I selects an incoming nucleotide according to its ability to form Watson–Crick-shaped pair with the template (Figure 3.4).
Polymerase I has three important functions; namely:
- 5′ → 3′ Polymerization activity,
- 3′ → 5′ Exonuclease activity and
- 5′ → 3′ Exonuclease activity.
If polymerase I erroneously incorporates a wrong nucleotide at the end of a growing chain, the polymerase activity is inhibited and 3′ → 5′ exonuclease activity excises the incorrect nucleotide. This is called ‘proofreading activity’.
Figure 3.4 Replication by DNA polymerase
Catalytic mechanism
Active site of DNA Pol-I has a shape complementary to the Watson–Crick base pairs. Although double-stranded DNA is mainly in the B conformation, the three base pairs near the active site assume the A conformation. The resulting wider and shallower minor groove permits protein side chains to form hydrogen bonds with the otherwise inaccessible N3 atoms of the purine bases and O2 atoms of the pyrimidine bases. DNA polymerases share a common catalytic mechanism for nucleotidyl transfer. Their active sites contain two metal ions usually Mg2+, which are ligated by two aspartate side chains. Metal ion B is liganded by all three phosphate groups of the bound dNTP, whereas metal ion A bridges the α-phosphate group of this dNTP and the primers 3′-OH. Metal ion A presumably activates the primer’s 3′-OH for a nucleophilic attack on the 5′-phosphate. Metal ion B function to orient its bound triphosphate group and to elctrostatically shield their negative charges as well as the additional negative charges on the transition state leading to the release of PPi. Polymerization rate 16–20 nucleotides/s. The processivity of the enzyme is 3–200.
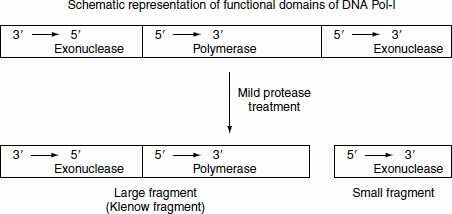
The 5′ → 3′ exonuclease activity of DNA Pol-I is located in a distinct structural domain of the enzyme and can be separated from the enzyme by mild protease treatment. When the 5′ → 3′ exo-nuclease activity is removed, the remaining fragment retains the polymerization activity and is called the large or ‘Klenow fragment’.
Functions of DNA Pol-I
- Physiologically the enzyme functions to repair DNA. Damaged DNA is endonucleolytically cleaved on the 5′ side of the lesion thereby activating the polymerase 5′ → 3′ exonuclease activity. While excising this damaged DNA, polymerase I simultaneously fills the resulting single-strand gap through its polymerase activity.
- Polymerase I catalyses ‘nick translation’. The polymerase I’s combined 5′ → 3′ exonuclease activity and polymerase activity can replace the nucleotides on the 5′ side of a singlestrand nick. These reactions in effect translate (move) the nick towards the DNA strands 3′-end without otherwise changing the DNA molecule. This nick translation is synthetically employed to prepare radiolabelled DNA.
- 3. Polymerase 5′ → 3′ exonuclease activity removes the RNA primer at the 5′-end of newly synthesized DNA while its polymerase activity fills in the resulting gap.
DNA Pol-II
The enzyme has a mass of 90 kDa.
- It has 5′ → 3′ polymerization activity.
- It has 3′ → 5′ exonuclease activity.
- Polymerization rate is about 40 nucleotides/s. This polymerase also fills the gaps and appears to facilitate DNA synthesis directed by damaged templates. Polymerase II has a low error rate but it is much too slow to be of any use in normal DNA synthesis. Polymerase II differs from polymerase I in that it lacks a 5′→3′ exonuclease activity (Table 3.1).
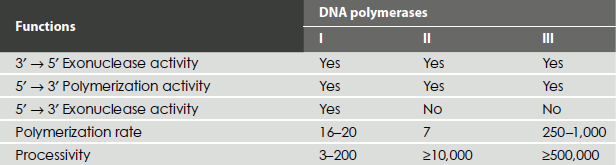
Table 3.1 Comparison of DNA polymerase of E. coli
DNA Pol-III
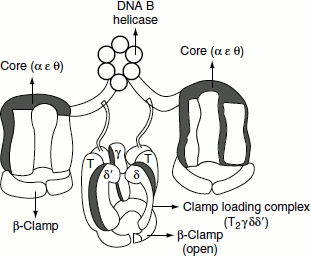
This enzyme is E. coli ‘DNA replicase’. The holoenzyme is a very large (>600 kDa) and highly complex protein composed of 10 different subunits. The core polymerase is composed of three subunits. The α-, ε- and the θ-subunits. The α-subunit contains the active site for nucleotide addition. The ε-subunit has 3′ → 5′ exonuclease activity and removes the incorrectly added nucleotides from the growing end. The θ is an accessory protein that stimulates the function of ε. Polymerase III holoenzyme consists of 10 subunits, which are α-, ε-, θ-, τ-, γ-, δ-, δ′-, χ-, ψ- and β-subunits. The β-subunit forms a donut-shaped dimer around the duplex DNA and holds the catalytic core polymerase near the 3′-end of the growing strand. Once tightly associated with the DNA, the β-subunit dimer functions like a ‘clamp’ that can slide freely along the DNA. In this way, the active site of core polymerase remains near the growing fork and the processivity of the enzyme is maximized. The five subunits of the enzyme; namely, γ, δ δ′,χ and ψ form the so-called γ complex that mediates two essential tasks: (Figure 3.5)
Figure 3.5 Bacterial DNA Polymerase III
- Loading of the β-subunit clamp on the duplex DNA—primer substrate, in a reaction that requires the hydrolysis of ATP.
- Unloading of the β-subunit clamp after the synthesis is completed. The τ-subunit acts to dimerize two core polymerases and is essential for co-ordinating the synthesis of the leading and lagging strands at each growing fork.
DNA Pol-V
Error prone synthesis occurs in E. coli. Functions involved in this pathway are identified by mutations in the genes umuD and umuC, which abolish UV-induced mutagenesis. This implies that umuD and umuC cause mutation to occur after UV irradiation. The genes constitute the umuDC operon and their expression is induced by DNA damage. Their products form a complex umuD′ 2C, consisting of two subunits of truncated umuD protein and one subunit of umuC. umuD′ 2C complex has DNA polymerase activity and is called DNA Pol-V and is responsible for synthesizing new DNA to replace sequences that have been damaged by UV. This is the only enzyme in E. coli that can bypass the pyrimidine dimers produced by UV.
Eukaryotic DNA Polymerases
There are many types of eukaryotic DNA polymerases; namely, DNA polymerases α, δ, ε, γ, β, ζ and η.
DNA polymerase α
It is a multi-subunit enzyme. It belongs to the A-family of DNA polymerases. It has primase activity. The larger subunit of the enzyme has polymerase activity. It does not have the proofreading activity. It is, therefore, unsuitable for high fidelity. DNA polymerase α is believed to function only in the synthesis of short primers. It has moderate processivity of about approximately 100 nucleotides.
Polymerase α/primase functions to synthesize 7–10-nucleotide-long RNA primers, which extend by an addition of approximately 15 nucleotides of DNA. Then, in a process called ‘polymerase switching’, replication factor C (RFC) displaces polymerase α and loads PCNA (proliferating cell nuclear antigen) on the template near the primer strand, following which polymerase δ binds to the PCNA and the processively extends the DNA.
DNA polymerase δ
It belongs to the B-family enzyme. It lacks primase and exhibits proofreading activity. It is a highly processive enzyme, but only when it is associated with PCNA. Polymerase δ, in association with PCNA, is required for both leading- and lagging-strand synthesis.
DNA polymerase ε
This also belongs to the B-family of nuclear enzymes. It superficially resembles polymerase δ. It has 3′ → 5′ exonuclease activity. The enzyme is highly processive even in the absence of PCNA.
DNA polymerase γ
It belongs to the A-family of enzyme. It occurs exclusively in the mitochondria. It replicates mitochondrial DNA. The chloroplast also contain similar enzyme.
DNA polymerase β
It belongs to the X-family of enzymes. It is involved in the base excision repair and the low-fidelity repair.
DNA polymerase ζ
It is involved in thymine dimer repair.
DNA polymerase η
It is involved in base damage repair (Table 3.2).
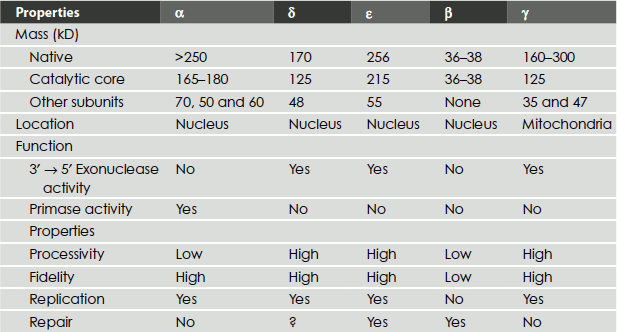
Table 3.2 Eukaryotic DNA polymerases
Helicases
Helicases are the enzymes that unwind nucleic acid molecules. There are DNA and RNA helicases. DNA helicases are essential during DNA replication, because they separate double-stranded DNA into single strands allowing each strand to be copied. As helicase unwinds the DNA, it forms the replication fork. The process of breaking the hydrogen bonds between the nucleotide bases pairs in double-stranded DNA requires energy from ATP or GTP hydrolysis. A helicase is generally multimeric. A common form of the helicase is a hexamer. This typically translocates along the DNA by using its multimeric structure to provide multiple DNA binding sites. Helicases are likely to have one conformation that binds the duplex DNA and another conformation that binds the single-stranded DNA. Alternation between the conformations drives the motor that melts the duplex and require ATP hydrolysis. Typically, one ATP is hydrolysed for each base pair that is unwound. Helicases may function with a particular polarity that is they can be either 3′→5′ or 5′→ 3′ helicase.
Helicases have been classified into five super families:
Superfamily I: It includes the helicases such as UvrD (E. coli, DNA repair), Rep (E. coli, DNA replication), PcrA (Staphylococcus aureus, recombination), Dda (bacteriophage T4, replication initiation) and RecD (E. coli, recombinational repair). RNA helicases also belong to this family. They play a role during viral RNA replication.
Superfamily II: It includes the enzymes such as rec Q (E. coli DNA repair).
Superfamily III: It consists of the helicases that are encoded mainly by the small DNA viruses and some large nucleocytoplasmic DNA viruses.
Superfamily IV, DnaB-like family: It includes the enzymes such as DnaB (E. coli, replication), gp41 (bacteriophage T4, DNA replication) and T7gp4 (bacteriophage T7, DNA replication).
Superfamily V, Rho-like family: It includes the enzymes such as Rho protein (E. coli, transcription termination).
Primases
Primase catalyses the formation of RNA primers required to initiate DNA replication. Primers are short RNA segments complementary to single-stranded DNA templates. Primase is of key importance in DNA replication because no known DNA polymerases can initiate the synthesis of a DNA strand without an initial RNA or DNA primer. The gene locus DnaG codes for the enzyme primase, an RNA polymerase in prokaryotes. In eukaryotes, the polymerase α has a primase subunit that synthesizes the primer.
Single-Strand Binding Proteins
Single-strand binding (SSB) binds the unwinded single strands of DNA, protecting it and preventing it from rewinding. E. coli SSB is a tetrameric molecule while eukaryotic SSB is trimer. Eukaryotic SSB is also called RPA. SSB binds the DNA co-operatively in which the binding of one monomer favours the binding of the other.
Ligases
DNA ligases close nicks in the phosphodiester backbone of DNA. Biologically, the DNA ligases are essential for the joining of Okazaki fragments formed during replication and for completing the DNA synthesis occurring in DNA repair process. There are two classes of DNA ligases. The first uses NAD+as a cofactor and only found in bacteria. The second uses ATP as a cofactor and found in eukaryotes, viruses and bacteriophages. The RNA primers in the lagging strand are removed by the 5′ → 3′ exonuclease activity of DNA Pol-I and replaced with DNA by the same enzyme. The resulting nick is sealed by DNA ligase (Figure 3.6).
Figure 3.6 Mechanism of action of DNA ligase
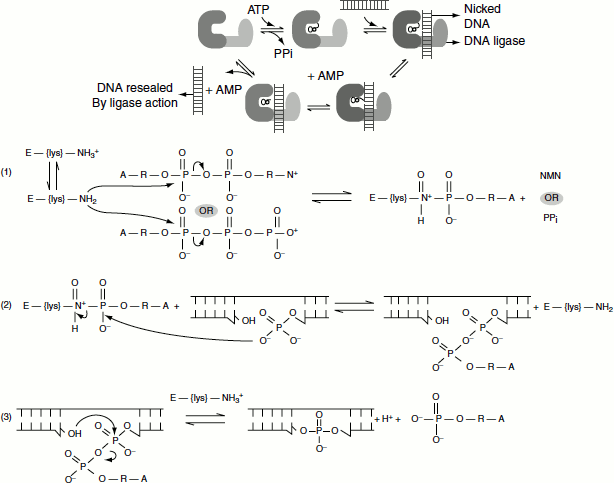
DNA ligase mechanism
The reaction occurs in three stages in all DNA ligases:
- Formation of a covalent enzyme-AMP intermediate linked to a lysine side-chain in the enzyme.
- Transfer of the AMP nucleotide to the 5′ -phosphate of the nicked DNA strand.
- Attack on the AMP-DNA bond by the 3′ -OH of the nicked DNA sealing the phosphate backbone and resealing AMP.
Figure 3.6 illustrates the three reaction stages:
When the energy source is ATP as in viruses and eukaryotes during adenylation of the enzyme, DNA ligase pyrophosphate is released. When the energy source is NAD+ as in prokaryotes, NMN is released.
Topoisomerases
In the cell, DNA is not free to rotate on its own axis. In E. coli, the closed circular chromosome clamps the DNA. In eukaryotes, DNA is arranged in fixed loops and is attached to several proteins; therefore, free rotation becomes impossible. However, the separation of DNA strand demands the rotation of the DNA duplex. This causes over winding, creating positive supercoils ahead of the replication fork and as the helix tightens further separation is resisted. If unrelieved, this tension halts strand separation and DNA replication. Topoisomerases catalyse the removal of the positive supercoils formed ahead of the replication fork. Topoisomerase change the topology of the DNA (discussed in chapter 1).