θ Replication
Autoradiography of replicating DNA was obtained by John Cairns. This indicates how DNA replicates. Autoradiogram of circular chromosome grown in medium containing 3H thymidine shows the presence of ‘replication’ ‘eyes’ or ‘bubbles’. These are called ‘Θ structures’, because of their resemblance to Greek letter theta. This indicates that double-stranded DNA replicates by the progressive separation of its two parental strands accompanied by the synthesis of their complementary strands to yield two semi-conservatively replicated duplex daughter strands. DNA replication involving Θ structures is known as ‘Θ replication’.
Rolling Circle Replication
It occurs during viral DNA replication in E. coli during mating. According to this model, by some initiation events, a nick is made in the duplex circle and this has 3′-OH and 5′-PO4 termini. Under the influence of a helicase and SSB protein, a replication fork is generated. The synthesis of primer is unnecessary because of the 3′-OH group. So that, leading-strand synthesis proceeds by elongation from this terminus. At the same time, the parental template for lagging-strand synthesis is displaced. Polymerase used for this synthesis is polymerase III. The displaced parental strand is replicated in the usual way by means of precursor fragments. The result of this mode of replication is a circle with a linear branch. It resembles the Greek letter sigma and is also called as sigma replication.
Bacteriophage ϕ×174 Replicates by Rolling Circle Replication
The virus carries a single-stranded DNA known as its (+) strand. On infecting an E. coli this strand directs the synthesis of its complementary strand or (−) strand to form the circular duplex replicative form (RF).
Synthesis of (−) strand
- The (+) strand is coated with SSB, except for a 44-nucleotide-long hairpin known as primosome assembly site (PAS). This is recognized and bound by PriA, PriB and PriC proteins.
- DnaB and DnaC proteins in the form of DnaB6 and DNAC6 complex add to the DNA with the help of DnaT protein in an ATP-requiring process. DnaC protein is then released yielding the pre-primosome. The pre-primosome in turn binds primase yielding the primosome.
- Primosome is propelled in the 5′ → 3′ direction along the (+) strand by PriA and DnaB helicase at the expense of ATP.
- Primase synthesizes an RNA primer.
- Polymerase III extends the primer to form the Okazaki fragments.
- Polymerase I excises the primer and replaces them by DNA. The fragments are then joined by DNA ligase and supercoiled by DNA gyrase to form ϕ174, (−) DNA.
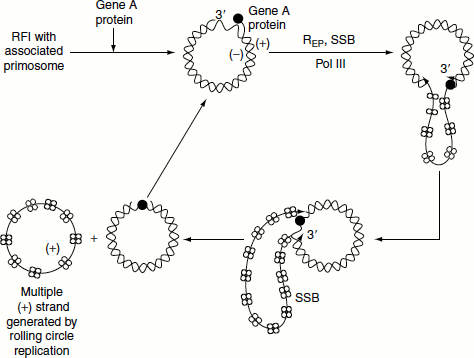
Synthesis of (+) strand by looped rolling circle replication
- (+) Strand synthesis begins with the primosome-aided binding of the gene A protein to its 30-bp recognition site.
- Gene A protein cleaves a specific phosphodiester bond on the (+) strand nucleotide by forming a covalent bond between a Tyr residue and the DNA’s 5′-phosphoryl group, thereby conserving the cleaved bond’s energy.
- Rep helicase subsequently attaches to the (−) strand at the gene A protein and with the aid of the primosome still associated with the (+) strand, commences the unwinding of the duplex DNA from (+) strand’s 5′-end.
- The displaced (+) strand is coated with SSB that prevents from reannealing to the (−) strand. E. coli polymerase III holoenzyme extends the (+) strand from its free 3′-OH group.
- Extension process generates a looped rolling circle structure in which the 5′-end of the old (+) strand remains linked to the gene A protein at the replication fork.
- It is thought that as the old (+) strand is peeled off the RF, the primosome synthesizes the primers required for the later generation of new (−) strand.
- When it has come full circle around the (−) strand, the gene A protein again makes a specific cut at the replication origin so as to form a covalent linkage with the new (+) strand 5′-end. Simultaneously, the newly formed 3′-OH group of the old and looped out (+) strand nucleophilically attacks its 5′-phosphoryl attachment to the gene A protein thereby liberating covalently closed (+) strand.
Synthesis of ϕ X174 (−) strand on a (+) strand template
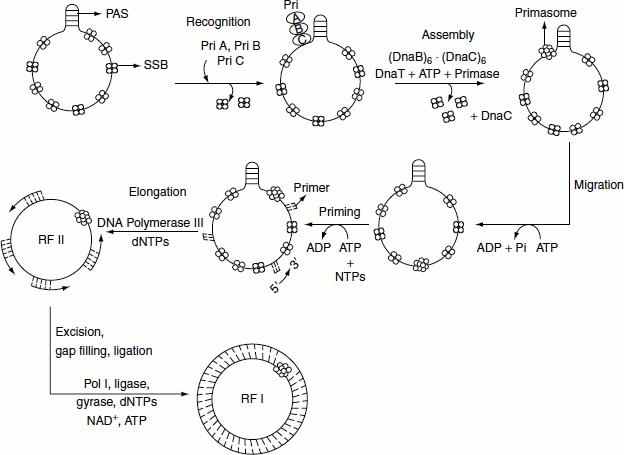
The synthesis of ϕ X174 (+) strand by rolling circle replication
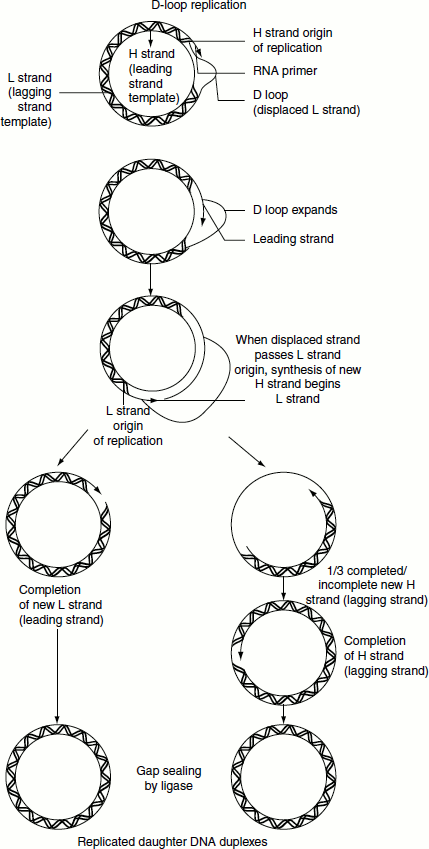
D-loop Replication
Mitochondrial and chloroplast DNA is replicated by a process in which leading-strand synthesis precedes lagging-strand synthesis. Leading strand, therefore, displaces the lagging-strand template to form a displacement or D-loop. The 15-Kb circular mitochondrial chromosome normally contains a single 500–600-nucleotide D-loop that undergoes frequent cycles of degradation and resynthesis. During replication, the D-loop is extended. When it has reached a point approximately two thirds of the way around the chromosome, the lagging-strand origin is exposed and its synthesis proceeds in the opposite direction around the chromosome. Lagging-strand synthesis is, therefore, only about one-third complete when the leading-strand synthesis terminates.