Whether a cell has only one chromosome or has many chromosomes, the entire genome must be replicated precisely once for ssevery cell division. Initiation of DNA replication commits the cell to further division. Once replication started, it continues until the entire genome has been duplicated. The unit of DNA in which an individual act of replication occurs is called the ‘replicon’. Each replicon fires only once in each cell cycle. The replicon is defined by its possession of the control elements needed for replication. It has an origin at which replication is initiated. It also has a terminus at which replication halts. The origin is a ‘cis -acting molecule’, i.e., it is able to affect only that molecule of DNA on which it resides. Bacteria and archaebacteria may contain additional genetic information in the form of ‘plasmids’. A plasmid is an autonomously circular DNA that constitutes a separate replicon.
The E. coli genome contains 4.7 × 106 nucleotide pairs. DNA replication proceeds at about 1,000 nucleotides/s and thus is done in not more than 40 min. One incorrect nucleotide is added for every 109 nucleotides inserted.
Origin of Replication
The replication starts at an origin by the separation of the two strands of the DNA duplex following a semi-conservative mode. When viewed under the electron microscope, the replicated region appears as a replication bubble. The point at which replication occurs is called replication fork also known as the growing point. A replication fork moves sequentially along the DNA from the origin. Replication may start either unidirectionally or bidirectionally from the origin. In E. coli, the origin is called oriC.
Initiation of DNA Replication
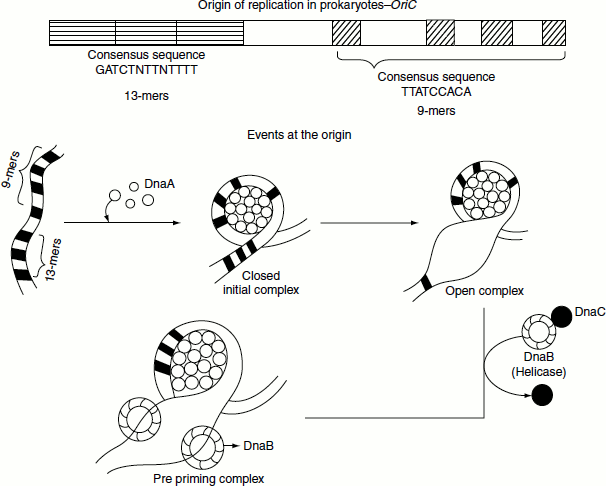
Events at the origin
E. coli oriC is a 240-bp DNA segment. oriC has a repetitive four 9-bp sequence and three AT-rich, 13-bp sequence referred to as ‘9-mers’ and ‘13-mers’ respectively. The 9-mer consensus sequence is 5′-TTATCCACA-3′. DnaA protein interacts with oriC to initiate replication. DnaA protein binds to the four 9-mers and causes it to become negatively supercoiled. Following this, the 13-mer sequences are melted. The 13-mer consensus sequence is 5′-GATCTNTTNTTTT-3′. Melting of the 13-mer sequences requires ATP that is hydrolysed by DnaA. Following melting, DnaA recruits hexameric helicase (six DnaB proteins) to opposite ends of the melted DNA. Recruitment of helicase requires six DnaC proteins, each of which is attached to one subunit of helicase. Once this complex is formed, an additional five DnaA proteins bind to the original five DnaA proteins to form five DnaA dimers. DnaC is then released and the ‘preprinting complex’ is complete. SSB protein is needed to prevent the single strands of DNA from forming any secondary structures and to prevent them from reannealing, and DNA gyrase is needed to relieve the stress (by creating negative supercoils) created by the action of DnaB helicase. The unwinding of DNA by DnaB helicase allows primase (DnaG) an RNA polymerase to prime each DNA template, so that DNA synthesis can begin. The DNA region at which all of the proteins come together to carry out the synthesis of daughter strand is called the replication fork, or ‘growing fork’. As replication proceeds, the growing fork and associated proteins move away from the origin. In order for DNA polymerase to move along and copy a duplex DNA, helicase must sequentially unwind the duplex and topoisomerase must remove the supercoils that form.
Regulation of replication initiation
In E. coli, DNA replication is regulated through several mechanisms, including the hemimethylation and sequestering of the origin sequence, the ratio of ATP to ADP and the levels of DnaA. All these control the process of initiator proteins binding to the origin sequences. E. coli methylates GATC DNA sequences. This results in the hemimethylated sequences that are recognized by the protein Seq A, which binds and sequesters the origin sequence. In addition, DnaA binds less to hemimethylated DNA. As the result, newly replicated origins are prevented from immediately initiating another round of replication. Elongation is a complex process involving many proteins.
Leading-and lagging-trand syntheses
A major complication in the operation of a DNA replication fork arises from two properties.
The two strands of parental DNA duplex are antiparallel and DNA polymerase can add nucleotides to the growing new strands only in the 5′ → 3′ direction.
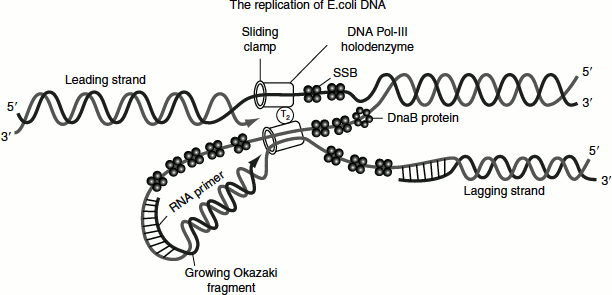
At the replication fork, the 3′ → 5′ parental DNA strand is copied by continuous synthesis initiated by RNA primer and proceeds in the direction of movement of the replication fork. Since this daughter DNA is continuously synthesized, it is called as the ‘leading strand’.
The 5′ → 3′ side of the parental strand is copied in the direction opposite to the movement of the replication fork by discontinuous synthesis. A cell accomplishes this by synthesizing a new primer every few hundred bases or so. Each of these primers base paired to the template is elongated in the 5′ → 3′ direction forming discontinuous segments called ‘Okazaki fragments’ after the discoverer Reiji Okazaki. The RNA primer of each Okazaki fragment is removed and replaced by DNA chain growth from the neighbouring Okazaki fragment. Finally, the enzyme DNA ligase joins the fragments. This discontinuously synthesized daughter strand is called the ‘lagging strand’.
DnaG primase binds to the DnaB helicase at the replication fork. DnaG primase initiates primer synthesis on the lagging strand followed by the binding of polymerase III. β-clamp is loaded by γ ‘clamp loader’ complex subunit onto the primer template junction. The polymerase III holoenzyme transits from the previously completed Okazaki fragment to the new primer terminus. DnaG primase is released and polymerase III synthesizes the new fragment. The τ processivity switch stops polymerase III when the end of previously synthesized Okazaki fragment reaches the active site of the enzyme.
The leading and lagging strands are synthesized concurrently
The co-ordinated synthesis of both leading and lagging DNA strands is thought to involve a dimeric DNA polymerase and a looping of the lagging strand, so that both strands can be synthesized in the same direction. A single molecule of polymerase III holoenzyme acts at the replication fork catalysing concurrently both leading- and lagging-strand syntheses. In prokaryotic systems, the directionality problem seen in lagging-strand synthesis is solved by the formation of a loop in the lagging strand at the replication fork to reorient the lagging-strand DNA polymerase, so that it advances in parallel with the leading-strand polymerase.
The two core-polymerase molecules at the fork are linked together by a τ-subunit dimer. The core polymerase synthesizing the leading strand moves, together with its β-subunit clamp, along its template in the direction of the movement of the fork, elongating the leading strand. The other core polymerase molecule that elongates the lagging strand moves with its β-subunit clamp in the direction opposite to that of the fork movement. As elongation of lagging strand proceeds, the size of the DNA loop between the core polymerase and the fork increases. The replication loop grows and shrinks during each cycle of the Okazaki fragment synthesis.
Double-stranded and newly synthesized DNA will be pushed into this loop. Eventually, the core polymerase synthesizing the lagging strand will complete an Okazaki fragment. It then dissociates from the DNA template, but the τ-subunit dimer continues to tether it to the fork protein complex. Simultaneously, DnaG primase binds adjacent to DnaB helicase on the lagging-strand template and initiates the synthesis of another RNA primer. This is followed by the binding of the P-clamp and the rebinding of the core polymerase. This polymerase molecule then proceeds to elongate the RNA primer to form another Okazaki fragment. As each Okazaki fragment nears completion, the RNA primer of the previous fragment is removed by the 5′ → 3′ exonuclease activity of DNA Pol-I. This enzyme also fills in the gaps between the lagging-strand fragments, which then are ligated together by DNA ligase. The E. coli enzyme uses NAD as a cofactor, while T4 DNA ligase (and other phage and eukaryotic ligases) uses ATP as a cofactor. Although the two core polymerase molecules are linked by the τ-subunit dimer, they are oriented in the opposite directions. Thus, the 3′ growing ends of both the leading and lagging strands are close together but offset from each other. The two core polymerases can add deoxyribonucleotides to the growing strands at the same time and rate, so that leading- and lagging-strand syntheses occur concurrently.
Topoisomerase, helicase and SSB are required continually throughout elongation to relive torsional stress, unwind the replication fork and to keep DNA single-stranded respectively.
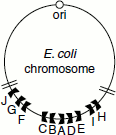
Figure 3.7 Termination sites of DNA replication in E. coli
Termination
Replication must be terminated properly both to dis-tangle the two daughter chromosomes and to regulate and co-ordinate replication with cell division.
The terminus of replication in E. coli is a large region of about 350 kb flanked by 10 nearly identical 23-bp terminator sites. These sites are non-palindromic and they are oriented differently on either side of the terminus. The Ter A, Ter D, Ter E, Ter I and Ter H are oriented in one direction, while Ter B, Ter C, Ter F, Ter G and Ter J are oriented in the other direction (Figure 3.7). They function in conjunction with a protein, Tus—terminator utilization substance, a 36-kDa protein, which depending on the orientation, permits a replisome to pass in one direction but not in the other. Tus interacts with DnaB and blocks its helicase activity, terminating replication (Figure 3.8).
Figure 3.8 Tus-Ter complex
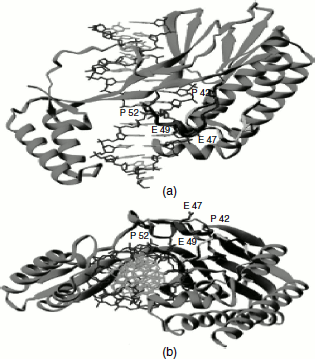
Newly replicated daughter chromosomes are prone to recombination reactions that may result in the production of circular dimers. These must be resolved into monomers before the segregation of the new DNA into two daughter cells. In E. coli, a specific locus called dif is required for resolution. dif is a 28-bp sequence located in the centre of the replication terminus region. Two recombination proteins, xer C and xer D, bind to the dif locus and catalyse a site-specific recombination reaction that generates two monomeric DNA. Another possible problem that arises at the termination of replication is the formation of catenated chromosomes. These are resolved by the concerted action of DNA gyrase and topoisomerase IV.
Fidelity of DNA Replication
The process of DNA replication is remarkably accurate. Error occurs once every 109–1,010 nucleotides incorporated. DNA polymerase, however, are not so accurate. They make mistakes once every 104–105 nucleotides incorporated. The proofreading activity of DNA polymerase improves the overall error rate by 102–103. However, this still leaves a difference of 102–103 in the error rates between DNA synthesis and replication. This difference is accommodated by mismatch repair systems that quickly fix any errors made during replication.