The eukaryotic transcription is more complex than the prokaryotic transcription due to the cellular compartmentalization. The genetic material in eukaryotes is primarily localized in the nucleus. A nuclear membrane separates the nucleus from the cytoplasm, where the protein synthesis takes place. The promoters for different genes are different. Each promoter contains a combination of sites to which specific protein factors bind. All of these factors help RNA polymerase to bind in the correct place and to initiate transcription. The three distinct RNA polymerases in the nucleus of a eukaryotic cell, defines the three major classes of the eukaryotic transcription unit.
The basal eukaryotic transcription complex includes the RNA polymerase and additional proteins that are necessary for the correct initiation and elongation. The eukaryotic RNA polymerases cannot find or bind to a promoter by themselves. Various assembly factors and positional factors help the RNA polymerase to locate the promoter and orient the polymerase correctly. The positional factor is the same in all cases.
Class-I Transcriptional Units
Class-I genes or transcriptional units are transcribed by the RNA Pol-I in the nucleolus. The best-studied examples are the rRNA transcription units. Each transcription unit consists of three rRNA genes—18S, 5.8S and 28S—and each unit is separated by a non-transcribed spacer. The eukaryotic nucleoli typically have many hundreds of copies of these transcription units that are tandemly arranged.
Promoters
RNA Pol-I has a bipartite promoter. The core promoter element spans positions from −31 to +6. The upstream control element (UCE) spans positions from −187 to −107. Both elements are closely related. There is approximately 85 per cent homology between them. These elements are unusual in that they are GC-rich.
Transcriptional complex
The eukaryotic RNA Pol-I does not contain the σ-factor, which can recognize the promoter and unwind the DNA double helix. In eukaryotes, these two functions are carried out by a set of proteins called ‘general transcription factors’.
Two additional transcription factors (the proteins that bind to the DNA and either repress or stimulate transcription) namely UBF1 and SL1 are known to be required to assist RNA Pol-I.
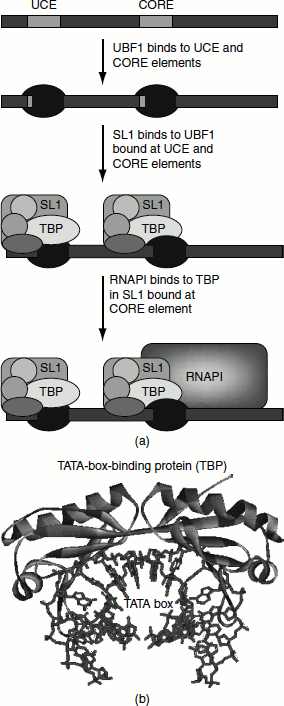
UBF1 is a single polypeptide that binds to the UCE and to the CORE promoter. It recognizes a GC-rich sequence within these elements and it is an assembly factor.
SL1 binds to the UBF1. It consists of four proteins, one of which is TATA-box-binding protein (TBP). TBP is required for the assembly of a transcriptional complex in all three classes of the eukaryotic transcription unit. TBP is a 180-amino acid protein that consists of two very similar 66-amino acid domains separated by a short basic region. The protein has a ‘saddle-shaped’ structure that sits astride a DNA molecule and binds to it via contacts in the minor groove. The binding also causes an 80° bend in the DNA (Figure 4.13 (b)).
SL1 is a positional factor—it positions the RNA polymerase at the promoter sequence, so that it initiates transcription in the correct place.
Once UBF1 and SL1 have formed a complex, RNA Pol-I binds to the CORE promoter to initiate transcription (Figure 4.13).
Figure 4.13 (a) Transcription by RNA polymerase I (b) Tata-box-binding protein (TBP)
Class-II Transcription Units
All genes that are transcribed and expressed via mRNA are transcribed by RNA Pol-II.
Promoters
The promoters used by RNA Pol-II have different structures depending upon the particular combination of transcription factors that are required to build a functional transcriptional complex at each promoter.
Some of the common class-II promoters are discussed in the following list.
The ‘TATA box’ [Goldberg-Hogness box] [−25 region] is located approximately 25 bp upstream of the start point of transcription. The consensus sequence of this element is TATAAAA (it resembles the TATAAT sequence of the prokaryotic −10 region). It is required for the correct positioning of the enzyme.
The ‘Initiator’ (Inr) is a sequence that is found in many promoters and defines the start point of transcription.
The ‘GC box’ is a common element in the eukaryotic class-II promoters. Its consensus sequence is GGGCGG. It may be present in one or more copies and can be located between 40 and 100 bp upstream of transcription start site. The transcription factor Sp1 binds to the GC box.
The ‘CAAT box’ is also often found between 40 and 100 bp upstream of the start point of transcription. Its consensus sequence is CCAAT. The transcription factor CTF or NF1 binds to the CAAT box.
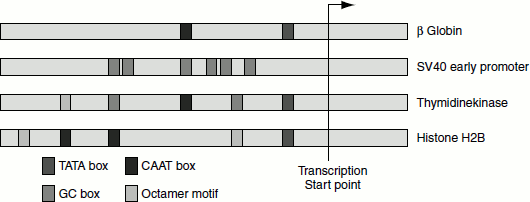
Figure 4.14 shows some examples of the eukaryotic promoters and the combination of sequence elements that they contain.
In addition to the above elements, enhancers may be required for full expression. They may provide an entry point for RNA polymerase or they may bind other proteins that assist RNA polymerase to bind to the promoter region.
Figure 4.14 Eukaryotic promoters
Enhancers and silencers
A typical protein-coding gene is likely to contain several enhancers that act at a distance. These elements are usually 700–1000 bp or more away from the start of transcription. Unlike the promoter elements, the enhancers can be downstream, upstream or within an intron; they can function in either orientation relative to the promoter. A typical enhancer is around 500 bp in length and contains the binding sites for several different transcription factors. An enhancer plays a role in the gene expression pattern. The enhancers increase the gene promoter activity either in all tissues or in a regulated manner (i.e., they can be tissue-specific or developmental stage-specific).
Similar elements that repress the gene activity are called ‘silencers’. The proteins bound at enhancer interact with the proteins that are bound at the promoter element most often through some intermediates called ‘co-activators’. Certain ‘transcriptional co-activators’ are ‘histone acetyltransferases (HATs)‘. Similarly, the transcriptional repression often requires both the binding of repressor on the silencer element and the participation of co-repressor proteins.
The binding of the activators to the enhancer element recruits HATs to relieve the association between histones and DNA, thereby it enhances the transcription.
The binding of the repressors to the silencer element recruits histone deacetylases (denoted by HDs or HDACs) to tighten the association between histones and DNA.
Insulators
The eukaryotic genomes are distinguished into the gene-rich euchromatin region and the gene-poor and highly condensed heterochromatin region. The heterochromatin region has a tendency to spread into neighbouring DNA. As a result, the natural barriers to spreading are required when the active genes are nearby. The ‘insulators’ are such transcriptional border that mediates the organization of transcriptional domains.
Figure 4.15 depicts the role of enhancers, silencer, activator repressor and coactivator.
Transcriptional complex
At least six ‘general (or basal) transcription factors’ (TFIIA, TFIIB, TFIID, TFIIE, TFIIF and TFIIH) have been characterized. In the presence of these transcription factors, the enzyme RNA Pol-II is able to initiate transcription at promoters correctly. However, even in the presence of transcription factors, the enzyme complex is unable to recognize and respond to regulatory signals.
Figure 4.15 Control elements of eukaryotic transcription
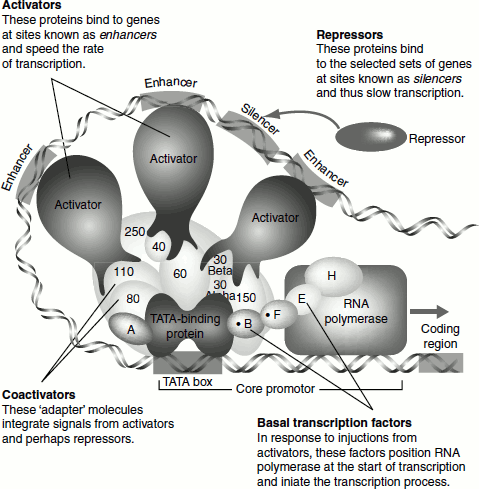
In addition to the general transcription factors, the transcriptional complex will also be affected by the presence of the promoter-proximal regulatory sequences and the presence of transcription factors that bind to those sequences. Such factors may be present in some cells/tissues but not in others. For example, the octamer motif (shown for the histone H2B gene above) binds two different transcription factors: Oct-1 and Oct-2. The Oct-1 is ubiquitous but the Oct-2 is expressed only in lymphoid cells where it activates k light chain gene transcription. Srb mediator, Srb-10-CDK and swi-Snf protein complexes confer the regulatory response capability on RNA Pol-II.
The transcription by RNA Pol-II occurs in four stages. They are:
I—Formation of initiation complex,
II—Initiation,
III— Elongation and
IV— Termination.
Formation of initiation complex
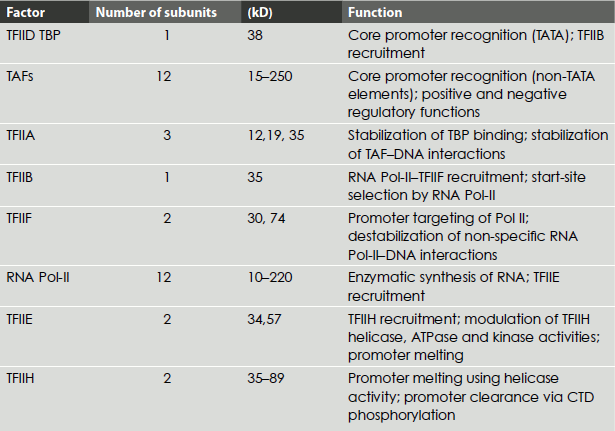
Initiation complex begins with the binding of the transcription factor TFIID to the TATA box. The TFIID is composed of one TATA-box-binding subunit called TBP. TFIIA then binds to form DA complex and activates TBP. TFIIB binds downstream of the TATA box, to DA complex, followed by the binding of a pre-formed complex between TFIIF and RNA Pol-II. Finally, TFIIE, TFIIH and TFIIJ must add to the complex for the initiation of the transcription. TFIIA, TFIIB and TFIID recognize the promoter-proximal DNA elements including the TATA box, the BRE (TFIIB recognition element), the Inr element and the downstream promoter element (DPE). The initiation by RNA Pol-II requires the energy derived from the hydrolysis of ATPs. One of the last factors to add to the complex is TFIIH, which has ATP-dependent helicase activity. TFIIH also has a protein kinase activity, which can transfer the γ-phosphates of ATPs to multiple serines in C-terminal repeat domain (CTD), the largest RNA Pol-II subunit. The helicase activity of TFIIH is also required for the nucleotide excision repair in the cell and the mutations in these subunits are associated with three genetic disorders: xeroderma pigmentosum, Cockayne’s disease (repair defects) and trichothiodystrophy (a transcription defect). The factor TFIIF also has an ATP-dependent helicase activity and could be involved in melting the DNA at initiation. TFIIE and TFIIH are required to melt DNA to allow polymerase movement (Figure 4.16) (Table 4.5).
Initiation
The helicase activity of the transcription factors results in the formation of an open promoter complex and exposes the template strand for transcription. RNA Pol-II contains two nucleotide-binding sites, which are the initiation site and the elongation site. The initiating nucleoside triphosphates bind to the enzyme and form hydrogen bond with a complementary base in DNA at the initiation site. The elongation site is then filled with nucleoside triphosphates. The two nucleotides are then joined together through a phosphodiester bond and the first base is released from the initiation site. RNA polymerase then moves, exactly by one nucleotide distance. After the initiation, TFIIE is released from the initiation complex.
Figure 4.16 Initiation of transcription by RNA pol-II – formation of the initiation complex
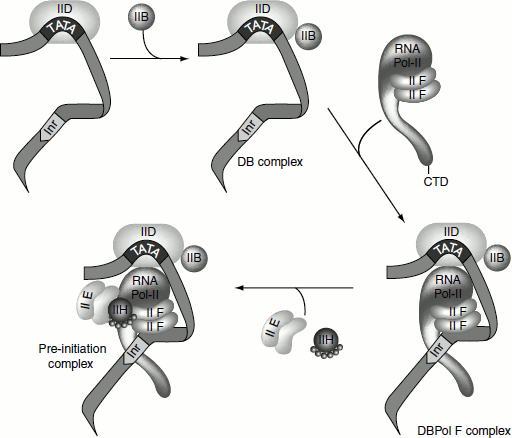
Table 4.5 Class II transcription initiation factors and their role
Elongation
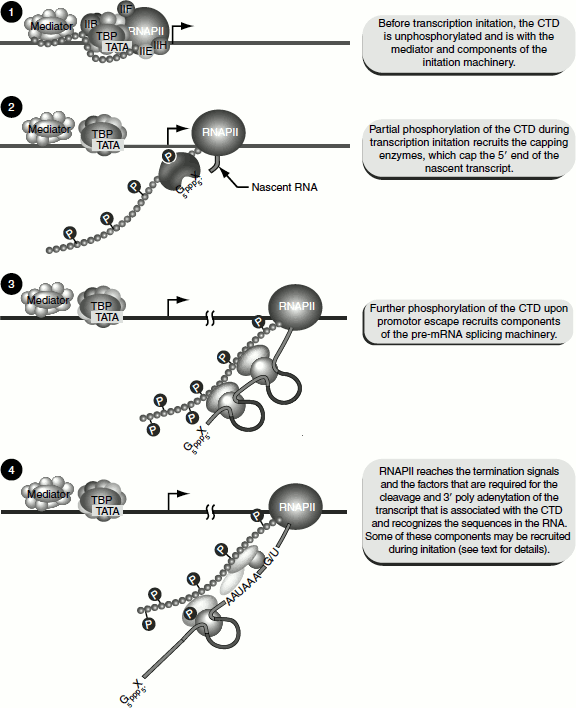
After the addition of a few nucleotides to the growing pre-mRNA chain, the protein kinase activity of TFIIH phosphorylates CTD of RNA Pol-II. This phosphorylation releases the attachment of CTD to TFIID and favours promoter clearance. RNA Pol-II can now move freely on DNA template. Most of the basal transcription factors are released at this stage.
Each site of phosphorylation on CTD serves as an anchor point for other proteins. The capping enzyme, guanyl transferase, which adds G residues to the 5′-end of the newly synthesized mRNA, binds to the CTD phosphorylated at serine 5. The serine 2 phosphorylation of the CTD is required for the splicing of the mRNA (Figure 4.17).
Figure 4.17 Promoter clearance and transcription elongation by RNA pol-II
RNA Pol-II unwinds DNA continuously, as the enzyme extends the growing RNA chain. Nascent RNA chain grows in the 5′→3′direction. The histone octamers are temporarily modified during the transit of RNA polymerase. The transcribed genes are repaired when a DNA damage occurs. TFIIH provides the link to a complex of repair enzymes.
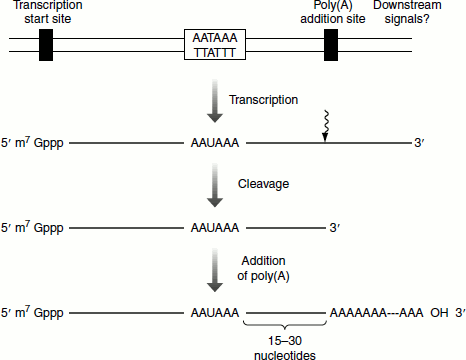
Figure 4.18 Eukaryotic transcription termination signalled by the AAUAAA sequence
Termination
Unlike in prokaryotes, the termination of eukaryotic transcription does not require rho factor. The end of the transcription of RNA Pol-II transcripts is signalled by a 3′-cleavage in the transcribed RNA. In most mRNA species, this is achieved by an upstream AAUAAA signal in co-ordination with downstream signals. Cleavage occurs normally about 15–30 nucleotides downstream of the AAUAAA element and AMP residues are subsequently added by poly(A) polymerase to form a poly(A) tail. Histone mRNA undergoes a different 3′-cleavage reaction (Figure 4.18).
Class-III Transcriptional Unit
Class-III transcriptional unit is primarily concerned with the transcription of tRNA, snRNAs (Small nuclear RNAs) and 5S rRNA genes and is transcribed by RNA Pol-III.
The promoters for 5S rRNA and tRNA genes are internal, i.e., they lie downstream of the start point. The promoters for snRNAs genes lie upstream. The promoters for RNA Pol-III may consist of bipartite sequences downstream of the start site with boxA separated from boxC (Type-1 promoter) or box BoxB (Type-2 promoter) or they may consist of separated sequences upstream of the start point. (Type-3 promoter).
Transcription of tRNA Gene
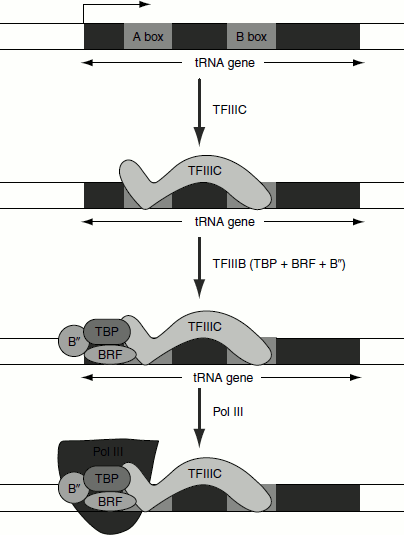
The tRNA gene contains Type-2 promoter, which consists of boxA sequence separated from a boxB sequence. The ‘pre-initiation complex’ (PIC) assembly begins with the binding of TFIIIC to both elements. First, TFIIIC, a large multi-subunit protein, binds with high affinity to the boxB promoter and with low affinity to the boxA. TFIIIC acts as an assembly factor and enables the binding of the trimeric positional factor TFIIIB. TFIIIB is made up of three subunits; the first is called TBP, the second is called BRF (TFIIB-related factor), which is similar in sequence to TFIIB, and the third subunit of TFIIIB, which is a 90-kD polypeptide and is called ‘B’. TFIIIB binding favours RNA polymerase binding and TFIIIC is released. RNA polymerase then initiates transcription in the presence of NTPs. After initiation, RNA polymerase elongates transcription and finally terminates transcription in a rhoindependent termination mechanism (Figure 4.19).
Figure 4.19 Transcription of tRNA gene by RNA pol III using type 2 promoter
Transcription of 5S rRNA Gene
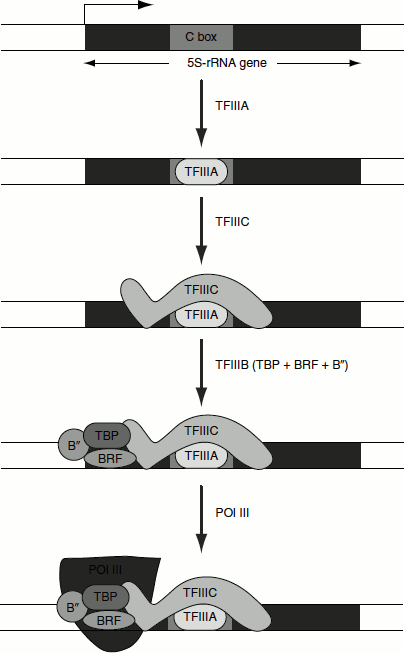
The 5S rRNA gene consists of type 1 promoters, which consists of boxA sequence separated by an intermediate element (IE) from boxC sequence. The entire boxA-IE-boxC sequence is referred to as the 5S internal control region (ICR). Internal type 1 promoter uses the assembly factors TFIIIA and TFIIIC to recruit the positioning factor TFIIIB, following which RNA polymerase binds with the release of TFIIIA and TFIIIC from the promoter, without affecting initiation reaction. TFIIIA has a zinc finger motif (discussed in Chapter 7). The efficiency of transcription is increased by the presence of the enhancer called promoter sequence element (PSE) and OCT (so called because it has an eight-bp-binding sequence) elements (Figure 4.20).
Figure 4.20 Transcription of 5S-rRNA gene by RNA pol III using type I promoter
RNA Pol-III elongates transcription and terminates it in a rho-dependent termination mechanism.
The snRNA genes have type-3 promoters and have three sequence elements that are all located upstream of the start point (OCT, PSE and TATA).