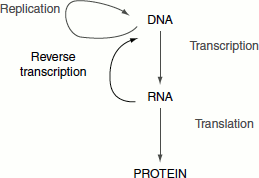
The discovery of reverse transcriptase has modified the central dogma of molecular biology, which held that genetic information should pass only from DNA to RNA. This enzyme synthesizes DNA from RNA, thus showing that the information can flow from RNA to DNA. The enzyme was discovered by David Baltimore and Howard M. Temin in the year 1970. Viruses such as human immunodeficiency virus (HIV), the Rous sarcoma virus, feline leukaemia virus and mouse mammary tumour viruses are some examples of RNA viruses containing reverse tran-scriptase. These viruses are called retroviruses, which are a group of animal viruses named for their backward (retro) way of replicating their nucleic acids. The unique feature of the retroviruses is their ability to produce DNA from RNA (Figure 4.36).
Figure 4.36 Reverse transcription
Reverse Transcriptase
The enzyme reverse transcriptase is RNA-directed DNA polymerase, which has three enzyme activities; namely,
- RNA-directed DNA polymerase,
- Ribonuclease H activity and
- DNA-directed DNA polymerase (Figure 4.37).
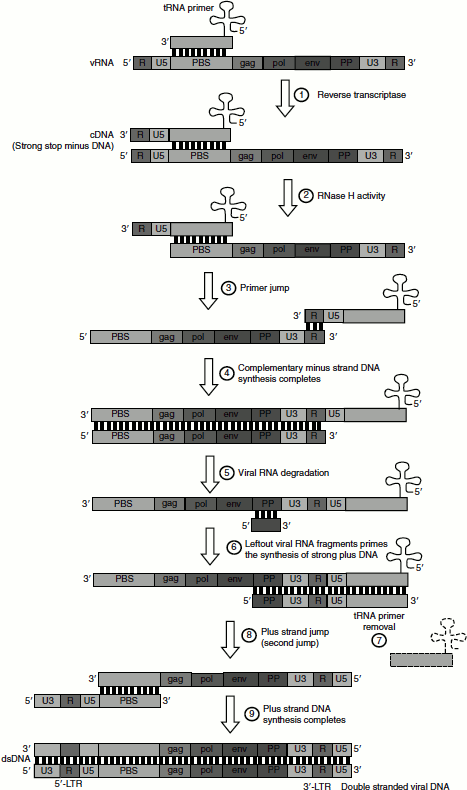
Figure 4.37 Mechanism of action of reverse transcripture
RNA-directed DNA polymerase
By using RNA-directed DNA polymerase activity, the reverse transcriptase produces a single-stranded DNA molecule using RNA as the template.
Ribonuclease H activity
H refers to hybrid; this activity frees the DNA from RNA-DNA hybrid.
DNA-directed DNA polymerase
dsDNA is synthesized from ssDNA. Reverse transcription is carried out by retroviruses, which utilize the host molecular machinery to replicate their RNA genetic material called the plus-strand RNA. The viral RNA has direct repeats at its ends. The sequence/direct repeats at the 5′-end is R-U5 and the sequence at the 3′-end U3R (long terminal repeats, LTRs).
Steps Involved in Reverse Transcription
- A specific cellular tRNA acts as a primer and hybridizes to a complementary part of the virus genome (plus-strand RNA) called the primer-binding site or PBS.
- Reverse transcriptase starts the synthesis of complementary minus-strand DNA.
- Enzyme reaches the end of the RNA template generating strong stop minus DNA.
- A domain on the reverse transcriptase enzyme called RNase H degrades the 5′-end of the RNA, which removes the U5 and R regions.
- The primer then ‘jumps’ to the 3′-end of the viral genome completing the synthesis of the complementary minus-strand DNA.
- tRNA primer is removed.
- Viral RNA is degraded, leaving fragments to prime the synthesis of DNA.
- Strong plus DNA is thus synthesized.
- Plus DNA is transferred to the other end of the minus strand in second jump.
- Plus-strand synthesis is completed.
- Integrase generates two base recessed 3′-ends in LTRs of the newly synthesized double-stranded DNA. It also generates staggered ends in host DNA.
- Integrase links recessed 3′-ends of LTR to staggered 5′-ends of the target.
- Thus, the RNA-directed DNA polymerization and subsequent integration into the host DNA help in the replication of the viral RNA using the host machinery.