The same secretory pathway is used by all eukaryotic cells for synthesizing and sorting secreted proteins and soluble luminal proteins in the ER, Golgi apparatus and lysosomes. These proteins are collectively referred to as ‘secretory proteins’.
Although all cells secrete a variety of proteins (e.g., extracellular matrix proteins), certain types of cells are specialized for the secretion of large amounts of specific proteins. For example, pancreatic acinar cells synthesize large quantities of digestive enzymes that are secreted into ductules that lead to the intestine.
The Signal Sequences of Secretory Proteins to ER
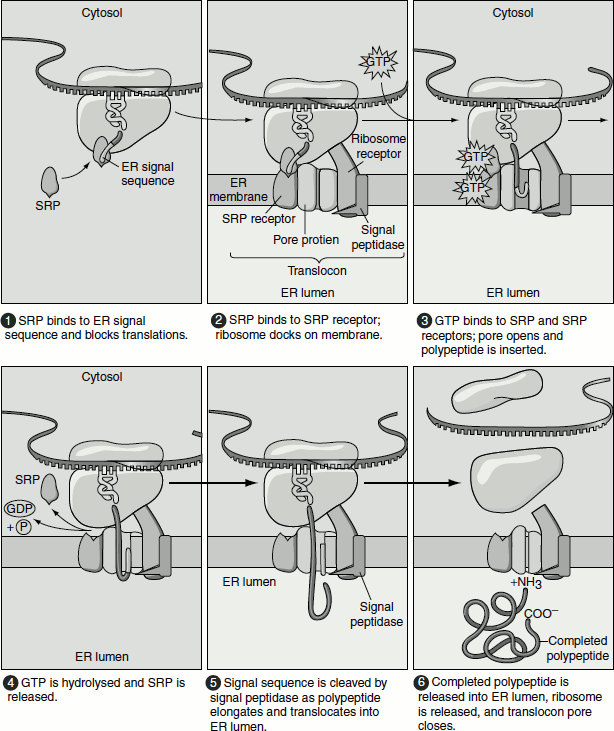
Soon after the synthesis of a secretory protein on free ribosomes in the cytosol starts, a 16–30-residue ER signal sequence in the nascent protein directs the ribosome to the ER membrane and initiates translocation of the growing polypeptide across the ER membrane (Figure 6.2). An ER signal sequence is located at the N-terminus of the protein and consequently is the first part of the protein to be synthesized.
The signal sequences of different secreted proteins contain one or more positively charged amino acids that are adjacent to a continuous stretch of 6–12 hydrophobic residues. The signal sequence of most secretory proteins is cleaved from the protein while it is still growing on the ribosome and thus is usually not present in the mature proteins that are found in cells.
The Signal Recognition Particle
Signal recognition particles (SRPs) are the key components in protein targeting. SRP is a cytosolic ribonucleoprotein particle that transiently binds simultaneously to the ER signal sequence in a nascent protein, to the large ribosomal unit and to the SRP receptor that are present on the membrane of the ER. Six discrete polypeptides and 300-nucleotide RNA compose the SRP. One of the SRP proteins P54 is chemically cross linked to the ER signal sequences. The hydrophobic region of P54 contains a cleft which interacts with the hydrophobic N-termini of nascent secretory proteins. This selectively targets them to the ER membrane. The SRP proteins P9 and P14 interact with the ribosome, while the SRP proteins P68 and P72 are required for protein translocation.
Figure 6.2 The SRP cycle-protein transport into the ER lumen
SRP Receptor
The SRP receptor is an integral membrane protein that is made up of two subunits α and β. Apart from mediating the interaction of nascent secretory protein with the ER membrane, the SRP receptor also permits the elongation and completion of the protein.
Thus, the SRP and SRP receptor function to bring ribosomes that are synthesizing secretory proteins to the ER membrane. The energy from GTP hydrolysis is used to release proteins lacking proper signal sequences from SRP and SRP receptor complex, thereby preventing their mis-targeting to the ER membrane.
The interaction of the SRP–nascent chain–ribosome complex with the SRP receptor is promoted when GTP is bound by both P54 subunit of SRP and the α-subunit of SRP receptor.
This is followed by the transfer of the nascent chain and ribosome to a site on the ER membrane where translocation can take place. Hydrolysis of the bound GTP takes place. After dissociating, SRP and its receptor release the bound GDP and recycle to the cytosol for initiating another round of interaction between ribosomes synthesizing nascent secretory proteins for their co-translational import to the ER.
Translocation into the ER Lumen
Co-translational translocation into ER
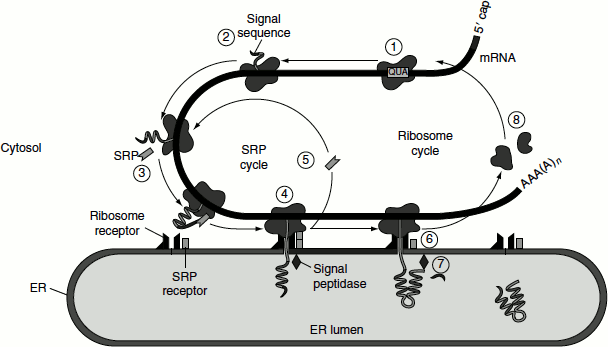
After the targeting of the ribosome-synthesizing secretory protein to the ER membrane, the ribosome and nascent chain are rapidly transferred to the ‘translocon’, a protein-lined channel within the membrane (Figure 6.3). The process of translation continues and the elongating polypeptide passes directly from the large ribosomal subunit into the central pore of the translocon. The 60S ribosomal subunit is aligned with the pore of the translocon. The growing chain is never exposed to the cytoplasm and does not fold until it reaches the ER lumen. To maintain the permeability barrier of the ER membrane, the translocon is regulated, so that it is open only when a ribosome–nascent chain complex is bound. Thus, the translocon is a gated channel. When the translocon first opens, a loop of the nascent chain, containing the signal sequence and approximately 30 adjacent amino acids can insert into the translocon pore. As the growing polypeptide chain enters the lumen of the ER, the signal sequence is cleaved by signal peptidase, which is a transmembrane ER protein associated with translocon. This protease recognizes a sequence on the C-terminal side of the hydrophobic core of the signal peptide and cleaves the chain specifically at this sequence once it has emerged into the ER lumen. The translocon remains open until translation is completed and the entire polypeptide chain has moved into the ER lumen.
Figure 6.3 Co-translational translocation into the ER lumen
Post-translational translocation into ER
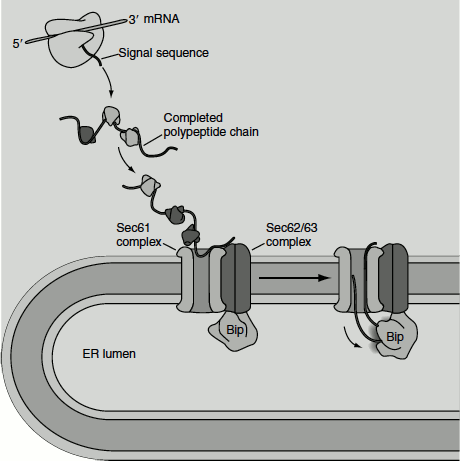
In most eukaryotes, secretory proteins enter the ER by co-translational translocation, using the energy derived from translocation to pass through the membrane. In yeast, however, some secretory proteins enter the ER lumen after translation has been completed, that is post-translational translocation (Figure 6.4). In this case, the translocating protein pass through the same translocon used in co-translational translocation; however, the SRP and SRP receptor are not involved in this case. In such cases, the direct interaction between the translocon and the signal sequence is sufficient for targeting to the ER membrane. In addition, the driving force for unidirectional translocation is provided by an additional protein complex known as the Sec63 complex and a member of the Hsc70 family of molecular chaperones known as Bip. The tetrameric Sec63 is embedded in the ER membrane in the vicinity of the translocon, while Bip is localized to the ER lumen. Bip has a peptide-binding domain and an ATPase domain. Bip binds and stabilizes the unfolded protein.
Figure 6.4 Post-translational protein import in to the ER
Once the N-terminal segment of the protein enters the ER lumen, signal peptidase cleaves the signal sequence. Bip–ATP interaction with the luminal portion of Sec63 complex causes the hydrolysis of the bound ATP, producing a conformational change in Bip that promotes its binding to an exposed polypeptide chain. In the absence of Bip, an unfolded polypeptide slides back within the translocon channel and thus does not allow the nascent polypeptide to enter the ER lumen. The Bip–ADP molecules bound to the polypeptide chain acts as a ratchet, ultimately drawing the entire polypeptide into the ER within a few seconds. Following this, the Bip molecules spontaneously exchange their bound ADP for ATP, leading to the release of the polypeptide, which can then fold into its native conformation. The recycled Bip–ATP is then ready for another interaction with Sec63.
Insertion of Proteins into the ER Membrane
Integral proteins located in ER, Golgi apparatus, lysosomal membranes and plasma membrane, which are synthesized on the rough ER that remain embedded in the membrane as they move to their final destinations along the same pathway followed by soluble secretory proteins. During this transport, the orientation of a membrane protein is preserved, i.e., the same segments of the protein always face the cytosol, while other segments always face the ER lumen. These sequences are collectively known as ‘topogenic sequences’, which direct the insertion and orientation of various classes of integral proteins into the membrane.
There are two major categories of hydrophobic signals used in the insertion of membrane proteins. All of these are membrane crossing domains:
- Start-transfer sequences: These are of two types:
- N-terminal signal peptide sequence: A cluster of about eight hydrophobic amino acids at the N-terminal end of a protein. This sequence remains in the membrane and is cleaved off of the protein after transfer through the membrane.
- Internal start-transfer sequence: Similar to a signal sequence, but located internally (not at the N-terminal end of the protein). It also binds to the SRP and initiates transfer. Unlike the N-terminal signal sequence, it is not cleaved after transfer of the protein.
- Stop-transfer signal: This is also a sequence of about eight hydrophobic amino acid residues. It follows either an N-terminal signal sequence or a start-transfer sequence. The stop-transfer signal is a membrane crossing domain. It remains in the membrane. The peptide is not cleaved.
- Start-transfer signal initiates the transfer of the carboxyl terminal arm of the polypeptide chain. When a stop-transfer peptide enters the translocator, it discharges the protein laterally into the membrane.
The topology of membrane proteins refers to the number of times that its polypeptide chains spans the membrane and the orientation of these membrane-spanning segments within the membrane. The key elements of a protein that determine its topology are the membrane-spanning segments themselves which usually contain 20–25 hydrophobic amino acids. Each of such segment forms an α-helix that spans the membrane with the hydrophobic amino acid anchored to the hydrophobic interior of the phospholipids bilayer. Based on the topogenic sequence, these integral transmembrane proteins are classified as ‘type I proteins’, when two sequences are involved in targeting and orienting them in the ER membrane, whereas ‘type II’ and ‘type III’ proteins contain a single, internal topogenic sequence.
Type I transmembrane protein insertion into the ER membrane
The signal sequence of all type I transmembrane proteins are located near their N-terminal. These proteins also have an internal hydrophobic sequence that becomes the transmembrane α-helix (transmembrane domain).
Like secretory protein, the N-terminal signal sequence on a nascent type I protein, initiates the co-translational translocation of the protein that is mediated through the combined action of the SRP and SRP receptor. Once the N-terminus of the growing polypeptide enters the lumen of the ER, the signal sequence is cleaved, and the growing chain continues to be extruded across the ER membrane.
A sequence of 22 hydrophobic amino acids in the middle of a type I protein stops the transfer of nascent chain through the translocon. This internal hydrophobic sequence can move laterally between the protein subunits that form the wall of the translocon. They get anchored in the phospholipid bilayer of the membrane, where it remains. This sequence that is responsible for the transmembrane anchoring of the protein is called a ‘stop-transfer anchor sequence’.
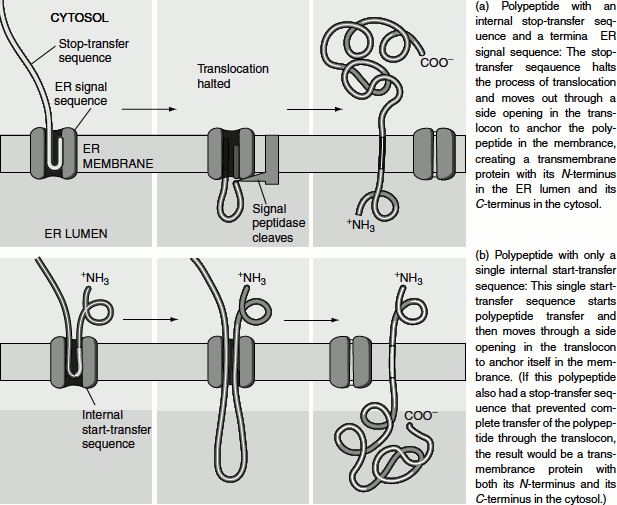
Once translocation is interrupted, translation continues at the ribosome, which is still anchored to the now unoccupied and closed translocon. As the C-terminus of the protein is synthesized, it loops out on the cytosolic side of the membrane. When translation is completed, the ribosome is released from the translocon and the C-terminus of the newly synthesized type I protein remains in the cytosol. That is the N-terminus region of the protein remains in the ER lumen, the hydrophobic region as the transmembrane domain and the C-terminal region as the cytosolic domain (Figure 6.5(a)).
Type II and type III transmembrane proteins insertion into the ER membrane
Unlike type I proteins, type II and type III proteins do not possess an N-terminal ER signal sequence. Instead they possess a single internal hydrophobic ‘signal-anchor sequence’ that functions both as an ER signal sequence and membrane anchor sequence. Based on the orientation of their respective signal anchor sequences within the translocon, type II and type III proteins have opposite orientation in the membrane.
In the case of type II proteins, after the internal signal anchor sequence is synthesized on a cytosolic ribosome, it is bound by an SRP. This directs the ribosome–nascent chain complex to the ER membrane. This is similar to targeting of soluble secretory proteins except that the hydrophobic signal sequence is not located at the N-terminus and is not subsequently cleaved. In the translocon, the N-terminal portion of the synthesizing protein is oriented towards the cytosol. As the chain is elongated and extruded into the lumen, the internal signal anchor moves laterally out the translocon. This hydrophobic sequence then anchors the polypeptide chain in the phospholipids bilayer.
Once protein synthesis is completed, the C-terminus of the polypeptide is released into the lumen and the ribosomal subunits are released into the cytosol. Thus, in this case, the N-terminus of the protein is oriented towards the cytosol and the C-terminus towards the ER lumen (Figure 6.5(b)).
In the case of type III proteins, the signal anchor sequence is located near the N-terminus, inserts the nascent chain into the ER membrane with its N-terminus facing the lumen, just the opposite of type II proteins. The signal sequence of type III proteins also prevent further extrusion of the nascent chain into the ER lumen, functioning as stop-transfer sequence. Translational elongation of the C-terminus in the cytosol continues and the hydrophobic sequence moving laterally between the translocon subunits anchor the polypeptide in the ER membrane.
Figure 6.5 (a) Type I transmembrane protein insertion into the ER membrane (b) Type II transmembrane protein insertion into the ER membrane
GPI-anchored Proteins
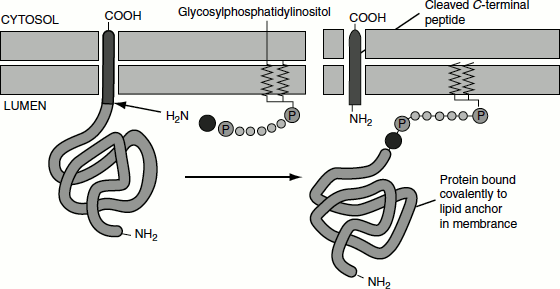
Some cell surface proteins are anchored to the phospholipid bilayer not by a sequence of hydrophobic amino acids but by a covalently attached amphipathic molecule, glycophosphatidylinositol (GPI). These proteins are synthesized and anchored to the ER membrane exactly like type I transmembrane proteins, with a cleaved N-terminal sequence and internal stop-transfer anchor sequence, directing the process. However, a short sequence of amino acids in the luminal domain, adjacent to the membrane-spanning domain, is recognized by a transamidase located within the ER membrane. This enzyme cleaves off the stop-transfer anchor sequence and transfers the remainder of the protein to a preformed GPI anchor in the membrane (Figure 6.6).
Figure 6.6 Membrane anchoring of proteins by GPI