Membrane and soluble secretory proteins that are synthesized on the rough ER undergo four principal modifications before they reach their final destination.
- Addition and processing of carbohydrates (glycosylation) in the ER and Golgi apparatus.
- Formation of disulphide bonds in the ER.
- Proper folding of polypeptide chains and assembly of multi-subunit proteins in the ER.
- Specific proteolytic cleavages in the ER, Golgi apparatus and secretory vesicles.
Protein Glycosylation
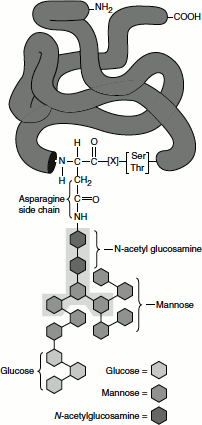
One or more carbohydrate chains are added to vast majority of proteins that are synthesized on the rough ER; indeed glycosylation is the principal chemical modification to most of these proteins. Carbohydrate chains in glycoproteins may be attached to the hydroxyl group in serine and threonine residues or to the amide nitrogen of asparagine. These are referred to as ‘O-linked oligosaccharides’ and ‘N-linked oligosaccharides’, respectively. O-linked oligosaccharides, such as those found in collagen and glycophorin, often contain only one to four sugar residues. The more common N-linked oligosaccharides are larger and more complex, containing several branches in mammalian cells. All N-linked oligosaccharides synthesis starts in the rough ER. A preformed oligosaccharide precursor containing 14 residues is attached to the protein. The branched oligosaccharide contains three glucose (Glc), nine mannose (Man) and two N-acetylglucosamine (GlcNAc)2 (Figure 6.7). This branched carbohydrate structure is modified in the ER and Golgi compartments, but five of the 14 residues are conserved in the structure of all N-linked oligosaccharides on secretory and membrane proteins.
The transfer of the oligosaccharide branch to the asparagine residue of the protein is catalysed by a membrane-bound enzyme, an oligosaccharyl transferase, which has its active site exposed on the luminal side of the ER membrane; this explains why cytosolic proteins are not glycosylated in this way. The precursor oligosaccharide is held in the ER membrane by a special lipid molecule called dolichol, and it is transferred to the target asparagine in a single enzymatic step immediately after that amino acid emerges in the ER lumen during protein translocation.
The oligosaccharide is assembled sugar by sugar onto the carrier lipid dolichol (a polyisoprenoid). Dolichol is long and very hydrophobic: its 22 five-carbon units can span the thickness of a lipid bilayer more than three times, so that the attached oligosaccharide is firmly anchored in the membrane. The first sugar group is linked to dolichol by a pyrophosphate bridge. This high-energy bond activates the oligosaccharide for its transfer from the lipid to an asparagine side chain of a nascent polypeptide on the luminal side of the rough ER. The synthesis of the oligosaccharide starts on the cytosolic side of the ER membrane and continues on the luminal face after the (Man)5 (GlcNAc)2 lipid intermediate is flipped across the bilayer. All of the subsequent glycosyl transfer reactions on the luminal side of the ER involve transfers from dolichol-P-glucose and dolichol-P-mannose; these activated and lipid-linked monosaccharides are synthesized from dolichol phosphate and UDP-glucose or GDP-mannose (as appropriate) on the cytosolic side of the ER and are then thought to be flipped across the ER membrane (Figure 6.8).
Figure 6.7 N-linked proteIn glycosylation
‘Tunicamycin’, an antibiotic produced by Streptomyces sp., mimics UDP-N-acetylglucosamine and blocks the first step in the synthesis of the core oligosaccharide of glycoproteins on dolichol phosphate. Tunicamycin group of antibiotics are produced by S. lysosuperficens. They contain uracil, N-acetylglucosamine, an 11-carbon aminodialdose called tunicamine, and a fatty acyl side chain. The structure of the fatty acyl side chain varies in the different members within the family. Apart from the variation in length of the fatty acyl side chain, some homologues lack the isopropyl group at the end and/or a 3-unsaturation.
Disulphide Bond Formation
Both intra-molecular and inter-molecular disulphide bonds (-S-S-) help stabilize the tertiary and quaternary structures of many proteins. The efficient formation of disulphide bonds in the lumen of the ER depends on the enzyme protein disulphide isomerase (PDI) which is present in all eukaryotic cells. This enzyme is especially abundant in the ER of secretory cells in the organs such as liver and pancreas.
Figure 6.8 Protein glycosylation in the ER lumen
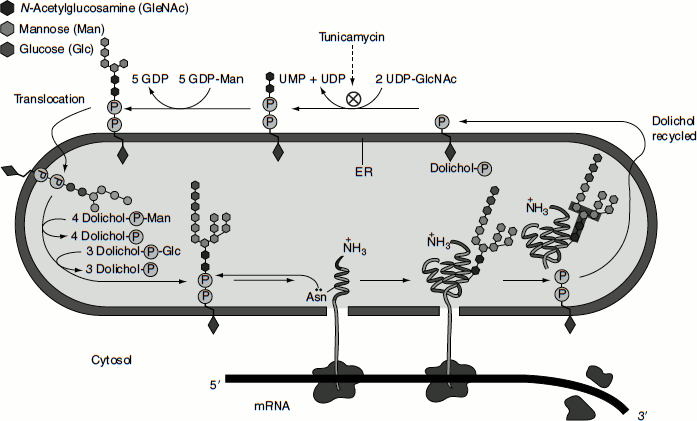
Figure 6.9 PDI-assisted disulphide bond formation
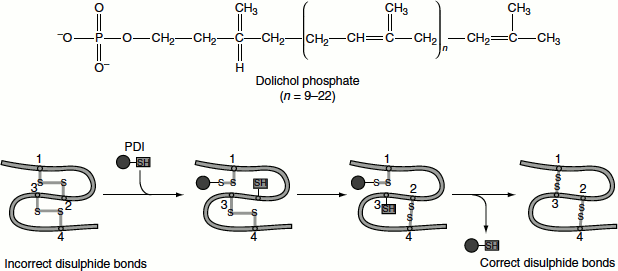
The disulphide bond in the active site of PDI can be readily transferred to a protein by two sequential thiol-disulphide transfer reactions. The reduced PDI generated by this reaction is returned to its oxidized form by the action of a protein called Ero1, which carries a disulphide bond that can be transferred to PDI (Figure 6.9).
The Role of N-linked Glycosylation in ER Protein Folding
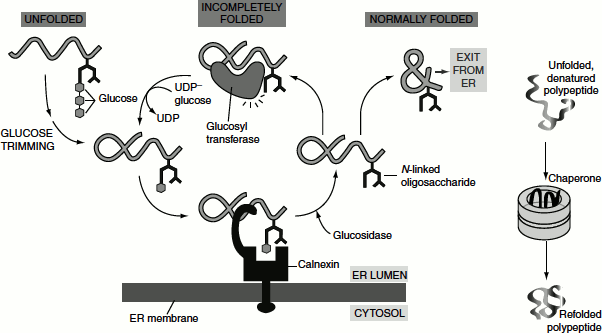
New soluble and membrane proteins produced in the ER generally fold into their proper conformation within minutes after their synthesis. The rapid folding of these proteins is mediated by the action of several proteins present in the ER called ‘chaperones’ (Figure 6.10).
The chaperone Bip not only helps co-translational translocation but also thought to prevent segments of a nascent chain from misfolding or forming aggregates, thereby promoting the folding of the polypeptide into proper conformation. PDI also contribute to proper folding.
The ER membrane-bound chaperone protein, ‘lectins’ (carbohydrate-binding protein) ‘calnexin’, binds to incompletely folded proteins containing one terminal glucose on N-linked oligosaccharides, trapping the protein in the ER. The removal of the terminal glucose by a glucosidase releases the protein from calnexin. A glucosyltransferase is the crucial enzyme that determines whether the protein is folded properly or not: if the protein is still incompletely folded, the enzyme transfers a new glucose from UDP-glucose to the N-linked oligosaccharide, renewing the protein’s affinity for calnexin and retaining it in the ER. The cycle repeats until the protein has folded completely. ‘Calreticulin’ functions similarly, except that it is a soluble ER resident protein. Another ER chaperone, ERp57, collaborates with calnexin and calreticulin in retaining an incompletely folded protein in the ER.
Other important protein folding catalyst in the ER lumen is ‘peptidyl prolyl isomerases’, a family of enzymes that accelerate rotation about peptidyl-prolyl bonds. Unfolded or misfolded proteins are often transported to the cytosol for degradation.
Figure 6.10 Protein folding in ER lumen