The nucleus is separated from the cytoplasm by two membranes which form the ‘nuclear envelope’. The nuclear membrane acts as barrier that prevent the free passage of molecules between the nucleus and the cytoplasm. The transport of macromolecules including mRNA, tRNAs and ribosomal subunits out of the nucleus and the transport of all nuclear proteins occur through the ‘nuclear pore’ (Figure 6.18). Numerous pores perforate the nuclear envelope in all eukaryotic cells. Each nuclear pore is formed from an elaborate structure termed the ‘nuclear pore complex’ (NPC). NPC is made up of multiple copies of some 50–100 different proteins called ‘nucleoporins’, which allow the regulated exchange of molecules between the nucleus and the cytoplasm. The selective traffic of proteins and RNAs through the NPCs not only establishes the internal composition of the nucleus, but also plays a critical role in regulating eukaryotic gene expression.
Figure 6.18 Nuclear transport and nuclear pore
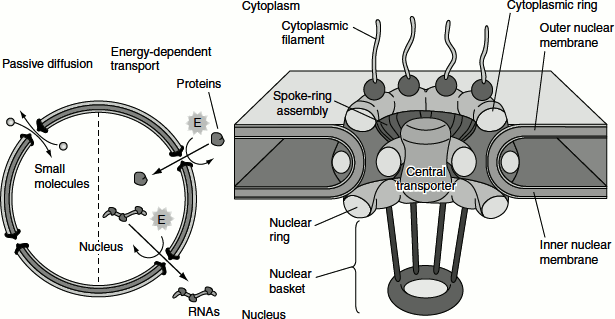
The nuclear membranes act as a barrier that separates the contents of the nucleus from the cytoplasm. Like other cell membranes, the nuclear membranes are phospholipid bilayers, which are permeable only to small non-polar molecules. The inner and outer nuclear membranes are joined at NPCs. These act as channels through which small polar molecules and macromolecules are able to travel through the nuclear envelope.
Small molecules and some proteins with molecular mass less than approximately 50 kDA pass freely across the nuclear envelope in either direction: cytoplasm to nucleus or nucleus to cytoplasm. They travel through open aqueous channels that are estimated to have diameters of approximately 9 nm, in the NPC. Most proteins and RNAs cannot pass through these open channels. These macromolecules pass through the NPC by an active process in which appropriate proteins and RNAs are recognized and selectively transported in only one direction (nucleus to cytoplasm or cytoplasm to nucleus).
The nuclear pore channels in the NPC, in response to appropriate signals, can open to a diameter of more than 25 nm. This size is sufficient to accommodate large ribonucleoprotein complexes, such as ribosomal subunits. Through these regulated channels, the nuclear proteins are selectively imported from the cytoplasm to the nucleus while RNAs are exported from the nucleus to the cytoplasm.
The NPC consists of an assembly of eight spokes attached to rings on the cytoplasmic and nuclear sides of the nuclear envelope. The spoke-ring assembly surrounds a central channel containing the central transporter.
Nuclear Localization Signals
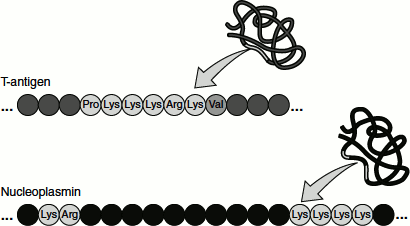
All proteins found in the nucleus are synthesized in the cytoplasm and imported into the nucleus through the NPC. Such proteins contain a ‘nuclear localization signal (NLS)’ that directs their selective transport into the nucleus (Figure 6.19). Most of these sequences, like that of T-antigen, are short stretches rich in basic amino acid residues (lysine and arginine). In many cases, however, the amino acids that form the NLS are close together but not immediately adjacent to each other. For example, the NLS of nucleoplasmin (a protein involved in chromatin assembly) consists of two parts: a Lys-Arg pair followed by four lysines located 10 amino acids farther downstream. Both the Lys–Arg and Lys–Lys–Lys–Lys sequences are required for nuclear targeting.
Figure 6.19 Nuclear localization signals
Transport of Proteins into the Nucleus
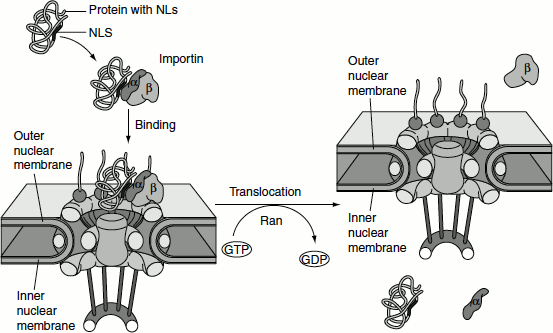
Protein import through the NPC can be divided into two steps, based on their requirement for energy (Figure 6.20). The first step does not require energy. In this step, proteins that contain NLS bind to the NPC but do not pass through the pore. The NLSs are recognized by a cytosolic receptor protein, and the receptor–substrate complex binds to the nuclear pore. ‘Karyopherins’ are a group of importin-β super-family proteins that are involved in transporting molecules through the pores of the nuclear envelope. Karyopherins may act as importins or exportins.
The receptor, called importin, consists of two subunits. One subunit (importin-α) binds to the basic amino acid-rich NLSs of proteins such as T-antigen and nucleoplasmin. The second subunit (importin-β) binds to the cytoplasmic filaments of the NPC, bringing the target protein to the nuclear pore. Other types of NLSs, such as those of ribosomal proteins, are recognized by distinct receptors which are related to importin-β and function similarly to importin-β during the transport of their target proteins into the nucleus.
The second step in nuclear import, translocation through the NPC, is an energy-dependent process that requires GTP hydrolysis. A GTP-binding protein called ‘Ran’, which is related to the Ras proteins, is involved in the process.
Enzymes that stimulate GTP binding to Ran are localized to the nuclear side of the nuclear envelope. Enzymes that stimulate GTP hydrolysis are localized to the cytoplasmic side. As the result, there is a gradient of Ran/GTP across the nuclear envelope, with a high concentration of Ran/GTP in the nucleus and a high concentration of Ran/GDP in the cytoplasm.
This gradient of Ran/GTP determines the directionality of nuclear transport. GTP hydrolysis by Ran provides the energy required for nuclear import. Importin-β forms a complex with importin-α and its associated target protein on the cytoplasmic side of the NPC, in the presence of a high concentration of Ran/GDP. This complex is then transported through the nuclear pore to the nucleus, where a high concentration of Ran/GTP is present. At the nuclear side of the pore, Ran/GTP binds to Importin-β, displacing importin-α and the target protein. As a result, the target protein is released within the nucleus. The Ran/GTP-Importin-β complex is then exported to the cytosol, where the bound GTP is hydrolysed to GDP, releasing Importin-β to participate in another cycle of nuclear import.
Figure 6.20 Nuclear import of proteins through the nuclear pore
The nuclear import of transcription factors is regulated directly by their phosphorylation. For example, the yeast transcription factor SWI5 is imported into the nucleus only at a specific stage of the cell cycle. Otherwise, SWI5 is retained in the cytoplasm as a result of phosphorylation at serine residues adjacent to its NLS, preventing nuclear import. Regulated dephosphorylation of these sites activates SWI5 at the appropriate stage of the cell cycle by permitting its translocation to the nucleus.
Transport of Proteins Out of the Nucleus
RNAs are transported across the nuclear envelope as RNA–protein complexes. These proteins are recognized by exportins and transported from the nucleus to the cytoplasm. Pre-mRNAs and mRNAs are associated with a set of at least 20 proteins (forming heterogeneous nuclear ribonucleoproteins (hnRNPs)) throughout their processing in the nucleus and eventually transport to the cytoplasm. At least two of these hnRNP proteins contain nuclear export signals and are thought to function as the carriers of mRNAs during their export to the cytoplasm; ribosomal RNAs are assembled with ribosomal proteins in the nucleolus and intact ribosomal subunits are then transported to the cytoplasm. Their export from the nucleus appears to be mediated by nuclear export signals present on ribosomal proteins. The tRNA must mature inside the nucleus before export. tRNAs are aminoacylated and only tRNAs charged with an amino acid are exported efficiently. Export occurs when the tRNA is carried through the nuclear pore by a complex of exportin-t and Ran/GTP. Exportin-t binds to tRNA which in turn complexes with Ran/GTP. This complex diffuse through the channel filled with FG proteins, which actually generate hydrophobic environment.
In contrast to mRNAs, tRNAs and rRNAs, which function in the cytoplasm, the snRNAs function within the nucleus as components of the RNA processing machinery. These RNAs are initially transported from the nucleus to the cytoplasm, where they associate with proteins to form functional snRNPs and then return to the nucleus. Proteins that bind to the 5′ caps of snRNAs appear to be involved in the export of the snRNAs to the cytoplasm, whereas sequences present on the snRNP proteins are responsible for the transport of snRNPs from the cytoplasm to the nucleus.
Transport and Sorting of Proteins to the Golgi Apparatus
The Golgi apparatus is sometimes referred to as ‘the post office of the cell’, as it processes proteins made by the ER and sends them out to their various destinations in the cell. Proteins enter the Golgi apparatus on the side that is facing the ER (cis side) and exit on the opposite side of the stack, which is facing the plasma membrane of the cell (trans side) (Figure 6.21).
Proteins make their way through the Golgi apparatus stack of intervening cisternae and along the way become modified. They are then packaged for transport to various locations within the cell. The Golgi apparatus cisternae vary in number, shape and organization in different cell types. There are three major cisternae (cis, medial and trans). Sometimes additional regions are added to either side, which are called the cis-Golgi network (CGN) and the trans-Golgi network (TGN). These networks have a more variable structure, including some cisterna-like regions and some vesiculated regions.
Different protein modification enzymes are present in each cisterna or the region of the Golgi apparatus. The Golgi enzymes catalyse the addition or removal of sugars from cargo proteins (glycosylation), the addition of sulphate groups (sulphation) and the addition of phosphate groups (phosphorylation). The enzymes sequentially add the appropriate modifications to the proteins.
Some Golgi-mediated modifications act as signals to direct the proteins to their final destinations that are present within cells; for example, the lysosome and the plasma membrane. Defects in various aspects of Golgi function can result in congenital glycosylation disorders, some forms of muscular dystrophy and may contribute to diabetes, cancer and cystic fibrosis.
Figure 6.21 Transport of proteins by the Golgi vesicles
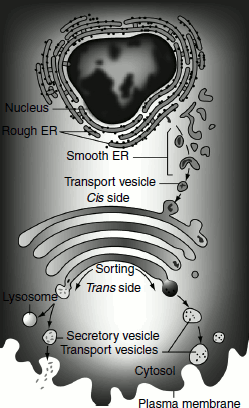
The Golgi apparatus is often found in close proximity to the ER in cells. Protein cargo moves from the ER to the Golgi apparatus, is modified vesicles to the Golgi apparatus, within the Golgi apparatus and is then sent to various destinations in the cell, including the lysosomes and the cell surface. Cargo proteins move between the Golgi cisternae by two possible explanations: the vesicular transport model and cisternal maturation model (Figure 6.22).
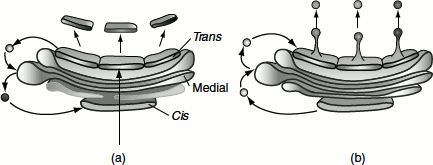
Figure 6.22 Two models of protein trafficking through the Golgi apparatus (a) The cisternal maturation model of protein movement through the Golgi apparatus. As a new cis cisterna is formed, it traverses the Golgi stack, changing as it matures by accumulating medial, then trans enzymes through vesicles that move from later to earlier cisternae (retrograde traffic). (b) The vesicular transport model, where each cisterna remains in one place with unchanging enzymes and the proteins move forward through the stack via vesicles that move from earlier to later cisternae (anterograde traffic)
Transport of Proteins to the Lysosomes
Proteins targeted to the lysosomes, for example enzymes such as hydrolases, upon arrival in the Golgi complex from the ER, their signal patch is recognized by a phosphotransferase that catalyses the phosphorylation of certain mannose residues in the enzymes’ oligosaccharides. The presence of one or more mannose-6-phosphate residues in their N-linked oligosaccharides is the structural signal that targets these proteins to lysosomes.
A receptor protein in the membrane of the Golgi complex recognizes this mannose-6-phosphate signal and binds the hydrolases so marked. Vesicles containing these receptor-hydrolase complexes bud from the trans side of the Golgi complex and make their way to sorting vesicles.
Inside the sorting vesicle, the receptor hydrolase complexes dissociate in a process facilitated by the lower pH within the sorting vesicles and by a phosphatase-catalysed removal of phosphate groups from the mannose-6-phosphate residues.
The receptor is then returned to the Golgi complex. Vesicles that are containing the hydrolases bud from the sorting vesicles and move to the lysosomes.
In cells that are treated with tunicamycin and hydrolases normally targeted for lysosomes do not reach their destination but are secreted instead, confirming that the N-linked oligosaccharide plays a key role in targeting these enzymes to lysosomes.