The tryptophan operon is required for the synthesis of the amino acid tryptophan. The trp operon is an example of a repressible operon—it is normally on; however, when a molecule called a ‘repressor’ is present, the operon turns off.
Structural Genes
The tryptophan operon contains five structural genes, namely trpE, trpD, trpC, trpB and trpA, that code for the enzymes involved in the synthesis of tryptophan. These genes are transcribed from a common promoter into a polycistronic mRNA, which is translated to yield the enzymes namely anthraanilate synthetase, indole-3-glycerol phosphate synthetase and tryptophan synthetase.
Regulatory Gene
The expression of the structural genes is controlled by a specific regulatory gene. The regulatory gene codes for a specific protein product called a repressor (sometimes called an apo-repressor). When the repressor is synthesized, it is inactive. However, it can be activated by complexing with the co-repressor (i.e., tryptophan). Thus, the trp repressor protein acts to reduce its own synthesis, i.e., it is ‘auto regulated’. The repressor is made as an inactive negative regulator. The amino acid tryptophan is co-repressor. When the levels of the amino acid tryptophan are high, two molecules bind to the dimeric trp repressor, changing its conformation to the active DNA-binding conformation and the repressor thus binds to the operator. This prevents RNA polymerase binding to the overlapping promoter sequence. Up to three trp repressor dimers can bind to the operator.
Operator
The active repressor/co-repressor complex acts by binding to a specific region of the DNA called the operator that is adjacent to the structural genes being regulated. Thus, in the presence of the co-repressor, the repressor is active and binds to the operator, resulting in the repression of the transcription of the structural genes. In contrast, in the absence of co-repressor, the repressor is inactive and does not bind to the operator, resulting in the transcription of the structural genes.
Co-repressor
The transcription of the tryptophan genes is influenced by the presence or the absence of a co-repressor (tryptophan).
The regulatory sites lie next to trpE and consist of a promoter site trpP, an overlapping operator site, trpO and a leader region trpL that codes for a leader peptide.
The enzymes that synthesize tryptophan are produced only when tryptophan is not available in the bacterial cell’s environment (medium).

trpE and trpD—They code for anthranilate synthetase.
trpC—It codes for indole-glycerol synthetase.
trpB and trpA—They code for tryptophan synthetase B chain and tryptophan synthetase A chain, respectively.
A complex system of repression and attenuation is used in the E. coli trp operon. The trp operon is a repressible operon with three transcription units. The first level of the control of gene expression is that the operon is negatively repressible that it is prevented from initiating transcription by its product, namely tryptophan. Attenuation is the second level of control, where the mRNA synthesis stops at a point about 140 nucleotides along the transcript. There is a region in the 5′ leader of the mRNA called the attenuator. An attenuator is an intrinsic terminator that is located between the promoter and the structural genes. Transcription is controlled by the rate of translation of the attenuator. High levels of tryptophan tRNA will attenuate or terminate transcription; on the other hand, low levels of tryptophan allow the trpEDCBA operon to be transcribed. This is made possible because of the changes in the secondary structure of the attenuator RNA, which are determined by the position of the ribosome on mRNA. The trpL gene codes for the ‘leader peptide’, which comprises 14 amino acids. The leader peptide comprises a ribosome-binding site whose AUG codon is followed by a short coding region that contains two successive codons for tryptophan.
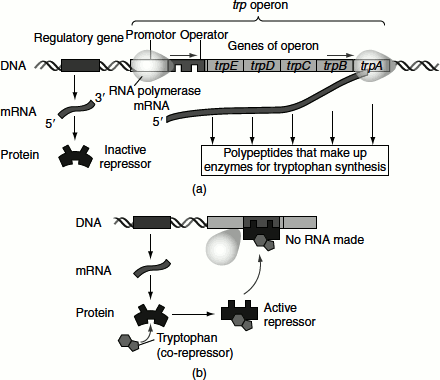
When the cell has low levels of tryptophan or in the absence of tryptophan, the repressor is inactive and does not bind to the operator. Transcription is initiated and is not terminated. Therefore, the structural gens are transcribed. As the result, a polycistronic mRNA is produced and this is translated to give the enzymes involved in tryptophan biosynthesis. Further, in the absence of trp tRNA, the ribosome stalls at the tryptophan codons and an alternative secondary structure formation prevents the formation of terminating hairpin structure and hence transcription continues (Figure 7.6 (a)).
On the other hand, when a cell has high levels of tryptophan, two molecules bind to the dimeric trp repressor, changing its conformation to the active DNA-binding conformation and the repressor thus binds to the operator. This prevents RNA polymerase binding to the overlapping promoter sequence inhibiting transcription. Further, in the presence of trp tRNA, the leader peptide is translated and the attenuator is able to form the hairpin that causes termination (Figure 7.6 (b)).
Figure 7.6 The tryptophan operon (a) Tryptophan absent, repressor inactive and operon on (b) Tryptophan present, repressor active and operon off
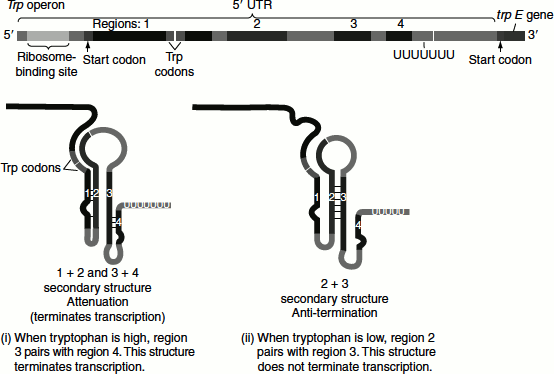
Attenuation
The trp operon attenuation mechanism uses signals encoded in four sequences namely sequences 1, 2, 3 and 4, within the leader region. The attenuator that is present at the end of the leader sequence is made up of sequences 3 and 4, which can base pair to form a G-C-rich stem and loop structure followed by a series of uridylate residues that resembles a transcription terminator. Transcription will halt here when this structure is formed (Figure 7.7).
The translation of the leader peptide begins immediately after it is transcribed and the bound ribosome follows closely behind the RNA polymerase as transcription proceeds.
When tryptophan levels are high, the concentration of charged trp tRNA is also high. Translation follows, the ribosome quickly translates sequence 1 and blocks sequence 2, before sequence 3 is transcribed by RNA polymerase. In this case, the sequence 2 is covered by the ribosome and this is unavailable for pairing with the sequence 3 when it is synthesized. The transcribed sequences 3 and 4 base pair, forming the terminator structure that halts the transcription of the structural genes.
When tryptophan levels are low, the ribosome pauses at the trp codons in sequence 1. Sequences 2 and 3 base pair with each other and this prevents the base pairing of the sequences 3 and 4 and consequently prevents attenuation. Thus, sequence 2 is an alternative complement for sequence 3. If sequences 2 and 3 base pair, the attenuator structure derived from the interaction of sequences 3 and 4 cannot form and transcription continues into the tryptophan biosynthetic genes.
This is an example of a ‘riboswitch’, a mechanism that can control transcription and translation through the interactions of molecules with an mRNA.
Figure 7.7 Attenuation and anti termination of tryptophan operon