Various molecular mechanisms operate to control gene expression at the level of transcription.
cis-Acting Regulatory Sequences: Promoters and Enhancers
These are the sequences that control the transcription of adjacent genes. Genes transcribed by RNA polymerase II have core promoter elements including the TATA box and Inr sequences. These cis-acting sequences serve as the binding sites of various transcription factors. Other cis-acting sequences serve as the binding sites of various regulatory factors that control the expression of individual genes. The cis-acting sequences are frequently located upstream of the TATA box; for example, consensus sequences such as CCAAT and GGGCGG (GC box).
In addition to the sequences mentioned above, many genes in mammalian cells are controlled by the regulatory sequences located farther away from the transcription start site. These sequences are called enhancers. The activity of the enhancers depends neither on their distance nor on their distance with respect to the transcription start site. They could stimulate the transcription when placed either upstream or downstream of the promoter, in either forward or backward orientation. Enhancers such as promoters function by binding transcription factors that regulate RNA polymerase. This is possible because of DNA looping, which allows a transcription factor bound to a distant enhancer to interact with proteins associated with RNA polymerase at the promoter.
The binding of specific transcriptional regulatory proteins to the enhancers is responsible for the control of gene expression during development and differentiation as well as during the response of cells to hormones and growth factors. An important feature of enhancers is that they usually contain multiple functional sequence elements that bind different transcriptional regulatory proteins. These proteins work together to regulate gene expression. The immunoglobulin heavy chain enhancer, for example, spans approximately 200 bp and contains at least nine distinct sequence elements that serve as protein-binding sites.
Though enhancers can act from considerable distance from theirs promoters, the activity of any given enhancer is specific for the promoter of its appropriate target gene. This activation is limited by boundaries called ‘insulators’ or ‘boundary elements’. Insulators define transcriptionally independent domains. They are also required to prevent the heterochromatin at centromeres and telomeres from spreading into euchromatin. That is insulators function to prevent the chromatin structure of one domain from spreading to its neighbours, thereby maintaining independently regulated regions of the genome.
Transcriptional Regulatory Proteins
A variety of regulatory proteins bind to promoter or enhancer sequences and regulate the gene expression.
Examples of transcription factors and their DNA-binding sites (Table 7.2).
Table 7.2 Transcription factors and their DNA-binding domains
| Transcription factors | DNA-binding sites |
|---|---|
| Specificity protein 1 (SP1) | GGGCGG |
| CCAAT/enhancer-binding protein (C/EBP) | CCAAT |
| Activator protein 1 (AP1) | TGACTCA |
| Octamer-binding protein (OCT-1 and OCT-2) | ATGCAAAT |
Eukaryotic Repressors
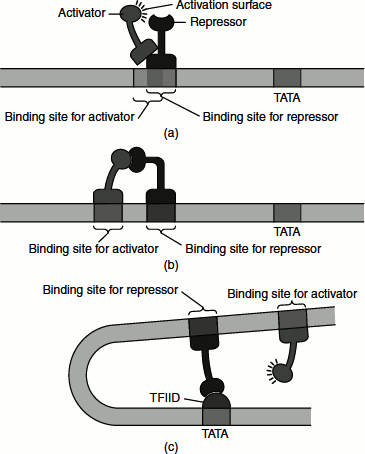
These bind to specific DNA sequences and inhibit transcription. In some cases, they interfere with the binding of other transcription factors to DNA. For example, the binding of the repressor near the transcription start site can block the interaction of RNA polymerase or general transcription factors with the promoter, which is similar to the action of repressors in bacteria. Some repressors compete with activators for binding to specific regulatory sequences. Certain repressors contain the same DNA-binding domain as that of the activator but lack its activation domain. Therefore, their binding to a promoter or enhancer blocks the binding of the activator, thereby inhibiting transcription initiation (Figure 7.11).
The functional targets of repressors are also diverse. Repressors act by interacting with specific activator proteins, with mediator proteins, with transcription factors and with ‘co-repressors’. One important role of repressors is to bring about the tissue-specific expression of genes in appropriate cell types; for example, repressor-binding site in the immunoglobulin enhancer is thought to contribute to its tissue-specific expression by suppressing transcription in non-lymphoid cell types. Other repressors play important roles in the control of cell proliferation and differentiation in response to hormones and growth factors.
RNA Interference
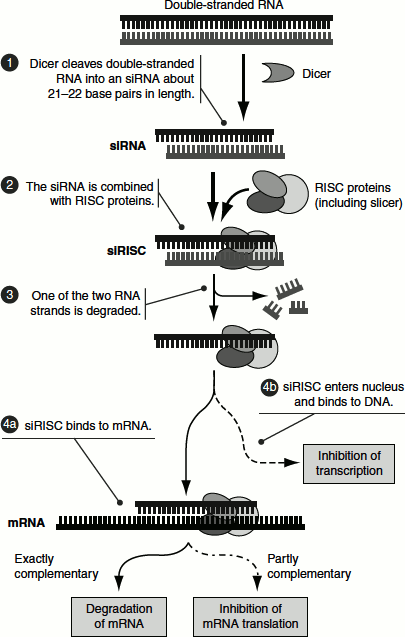
Short RNAs that have complementary sequence to the mRNA can silence gene expression. A complex of a double-stranded RNA is cleaved into short fragments of 21–22 bp in length by the ribonuclease.
Figure 7.11 Different modes of transcription repression in eukaryotes by repressors (a) Competitive binding with activator (b) Interaction with activation domain of bound activator (c) Interaction with general transcription factors
These fragments are called short interfering RNAs (siRNAs). The siRNAs bind to the RNA-induced silencing complex (RISC). One of the strands of siRNAs is degraded. The remaining single-stranded siRNA that is complexed with the RISC can then bind to complementary mRNA and the paired mRNA is cleaved. Further, the RISC–siRNA complex can enter the nucleus, bind to the genomic sequence and initiate a DNA methylation-based chromatin condensation and thus cause the inactivation of the gene (Figure 7.12).
microRNAs (miRNAs)
These are gene products that are 21–22 nucleotides in length. The primary miRNAs transcribed form hairpin structures. They are cleaved to make precursor miRNAs (roughly 70 nucleotides in length). They are then exported to the cytoplasm where they are further cleaved by enzymes into the 21–22 nucleotide mature miRNAs. The miRNAs form ribonucleoprotein complexes with mRNAs. If the match is exact, the mRNA is destroyed, similar to siRNA mechanisms (Figure 7.13).
Figure 7.12 Gene regulation by siRNAs