Regulatory proteins generally bind to specific DNA sequences. Their affinity for these target sequences is approximately 104–106 times higher than their affinity for many other DNA sequences. Most regulatory proteins have discrete DNA-binding domains containing substructures that interact closely and specifically with the DNA.
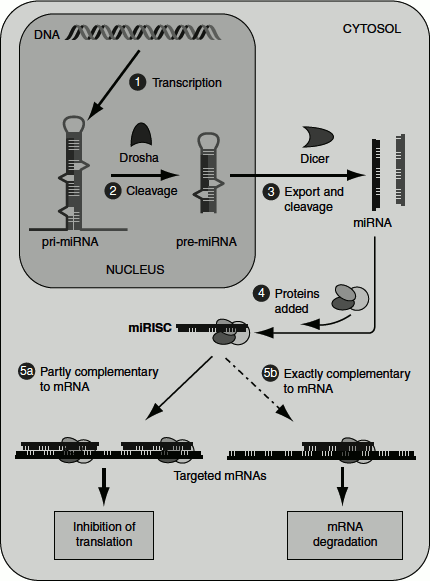
Figure 7.13 Gene regulation by miRNAs
Within the regulatory proteins, the amino acid side chains form hydrogen bonds to bases in the DNA. The amino acids mostly involved in such interactions include Asn, Gln, Lys and Arg. These mostly interact with the major groove of the DNA. This interaction requires a relatively small structure that can protrude from the protein surface. The DNA-binding domains of regulatory proteins tend to be small usually 60–90 amino acids in length. The DNA-binding sites for regulatory proteins are often the inverted repeats of short DNA sequence (a palindrome) at which multiple subunits of regulatory protein bind co-operatively.
Several DNA-binding motifs have been described of which two play prominent role in the binding of DNA by regulatory proteins; the helix-turn-helix (HTH) and the zinc finger.
Helix-Turn-Helix Motif
A ‘motif’ is a short conserved sequence pattern associated with some distinct functions of a protein. Structurally, a motif is a simple combination of a few secondary structure elements.
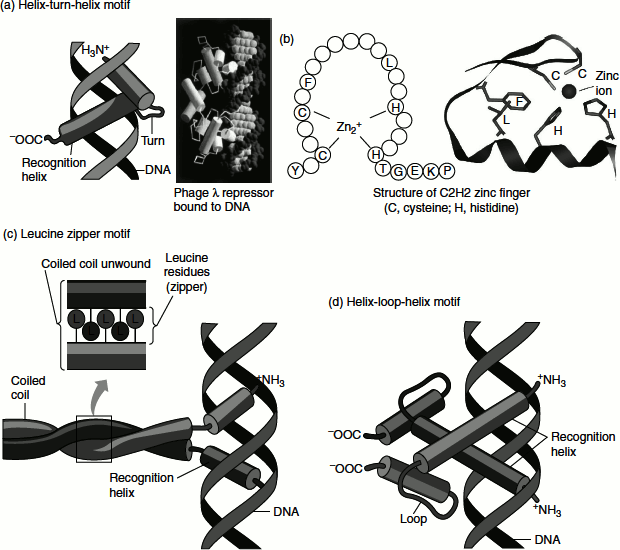
HTH motif—DNA-binding motif—is crucial to the interaction of regulatory proteins with the DNA. The HTH motif comprises about 20 amino acids in two a-helical segments, each is 7–9 amino acids long, separated by a β-turn. The two α-helical segments occupy the N-terminal and C-terminal ends of the motif and play a role in recognition and binding to DNA. This motif interacts with the major groove of the DNA through hydrogen bonds and Van der Waals’ interactions. The lac repressor has this DNA-binding motif (Figure 7.15 (a)).
Zinc-Finger Motif
Zinc finger consists of about 30 amino acids forming an elongated loop held together at the base by a single Zn2+ ion, which is co-ordinated to four amino acid residues (four Cys or two Cys and two His). The Zn2+ ion does not itself interact with DNA, rather the co-ordination of zinc with the amino acid residues stabilizes this small structural motif. Several hydrophobic side chains in the core of the structure helps to stabilize the structure (Figure 7.14) and (Figure 7.15 (b)). Many eukaryotic DNA-binding proteins contain zinc finger. The interaction of a single zinc finger with DNA is typically weak. Many DNA-binding proteins such as Zif 268 have multiple zinc fingers that substantially enhance the binding by simultaneously interacting with the DNA. Zinc fingers also function as RNA-binding motifs; for example, proteins that bind eukaryotic mRNAs, transcriptional repressors, etc.
Figure 7.14 DNA binding motifs
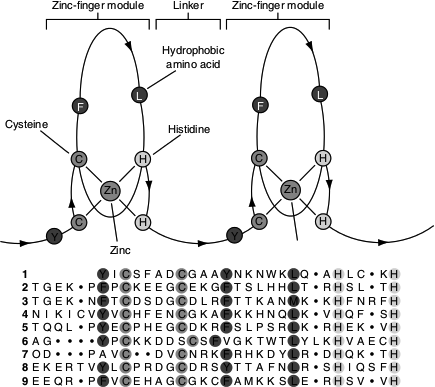
Figure 7.15 Protein-protein interaction domains
The Cys2His2-like fold group is by far the best-characterized class of zinc fingers and are extremely common in mammalian transcription factors; for example, transcription factor TFIIIA. Certain hormone receptors that bind DNA sequences also have zinc-finger motif; for example, the glucocorticoid receptor.
Homeodomain
This domain is made up of 60 amino acids. It is called as homeodomain because it was discovered in homeotic genes (the genes that regulate the development of body patterns). It is highly conserved and has been identified in a wide variety of proteins. The DNA-binding segment of the domain is related to the HTH motif. The DNA sequence that encode this domain is known as ‘homeobox’.
Figure 7.16 Homeo domain (a) Homeotic gene (b) Homeotic protein bound to DNA
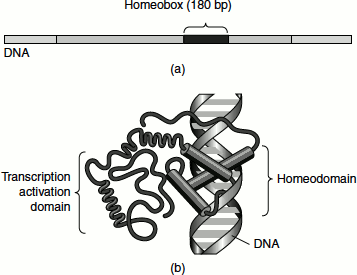
Homeodomain is 180 bp long. It binds to DNA in a sequence-specific manner. The homeodomain contains three well-defined alpha helices and a more flexible fourth helix. Helices 2 and 3 form a HTH motif. Helix 3 acts as DNA recognition site and interacts directly with the major groove of DNA. The flexible amino terminus of homeodomain establishes contact with the minor groove of DNA (Figure 7.16).