The integrity of the DNA is very much important for the sustainability of the cell. For this reason, a vast number of repair systems work efficiently debugging the errors. The repair process begins during replication and continues in various forms even during the post-replication. About 130 repair genes exist in human genome that codes for various proteins involved in DNA repair. DNA repair systems can be divided into the following types:
- Direct repair systems.
- Excision repair system, which include base excision repair (BER) and nucleotide excision repair (NER).
- Recombination repair systems.
- Error-prone repair—SOS repair systems.
Direct Repair Systems
Direct repair is rare and involves the reserval of the DNA damage; for example, ‘photoreactivation’ of pyrimidine dimers. These dimers are formed in the DNA by UV light. Enzymes called ‘photolyases’ are activated by visible light (300–600 nm) and they cleave the dimers to yield intact pyrimidines (Figure 8.9). The enzyme uses base flipping that reverses the bond between the pyrimidine dimers. The pyrimidine dimmer is flipped out of the double helix, into a cavity in the enzyme. Close to this cavity is an active site that contains an electron donor, which provides the electrons to break the bonds of the dimer. Energy required for the reaction is provided by visible light (Figure 8.10).
Another example for direct reversal is the action of methylguanine methyltransferase (MGMT). The enzyme recognizes O6 methylguanine in DNA and removes the methyl group. Such methyl groups are added to the bases by the action of alkylating agents such as methylmethane sulphonate (MMS). The enzyme removes the methyl group by attaching it to its own cysteine residues. This reaction restores the guanine but inactivates the enzyme. That is the entire protein molecule is expended for each O6 methylguanine that is being repaired (Figure 8.11).
Figure 8.9 Photoreactivation of Pyr-Pyr dimers
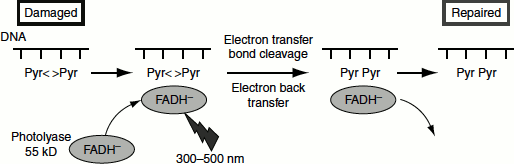
Figure 8.10 Photoreactivation
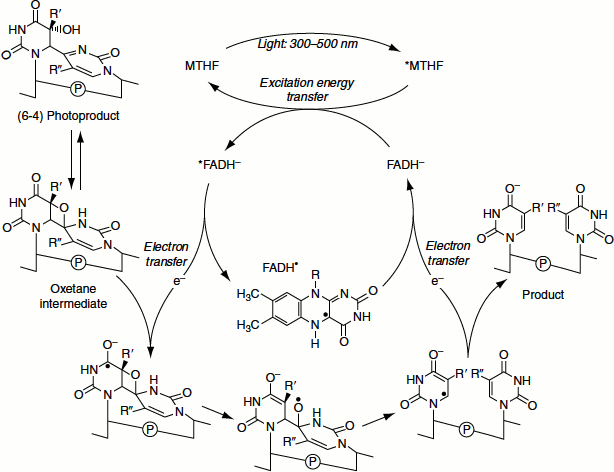
Figure 8.11 Direct reversal of methylation by MGMT
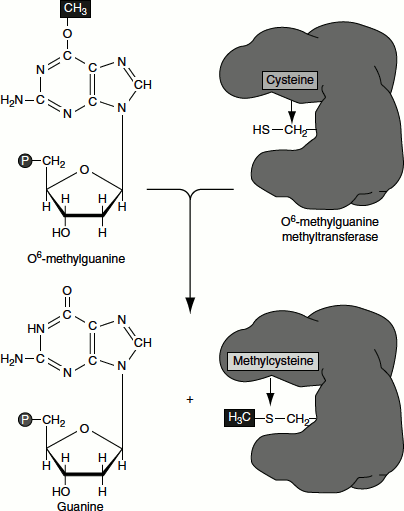
Excision Repair Systems
Excision repair deals with a variety of structural defects in the DNA. Mismatches between the strands of DNA are one of the major targets for repair systems. Mismatches are usually corrected by excision repair. There are two types of excision repair:
- Base excision repair (BER) and
- Nucleotide excision repair (NER).
In general, excision repair comprises two steps namely:
- Incision step: In this step, the damaged structure is recognized by an endonuclease that cleaves the DNA strand on both sides of the damage.
- Excision step: In this step, a 5′ → 3′ exonuclease removes a stretch of the damaged strand. Alternatively, a helicase displaces the damaged strand, which is subsequently degraded.
Base Excision Repair
This repair involves the direct removal of the damaged base from the DNA. This serves as the trigger to activate the enzymes that excise and replace a stretch of DNA, including the damaged site. Enzymes that remove bases from DNA are called glycosylases and lyases. Glycosylases cleave the bond between the damaged or the mismatched base and deoxyribose resulting in the formation of an ‘AP site or abasic site’ (apurinic or apyrimidinic site). Lyases acts similarly but takes the reaction further by opening the deoxyribose sugar ring using NH2 group.
Glycosylase action is followed by the action of endonuclease APE1, which cleaves the polynucleotide chain on the 5′ side. This in turn attracts the replication complex polymerase δ/ε and other ancillary components. A short DNA strand is synthesized extending for two to ten nucleotides. The displaced DNA material is removed by endonucleaseFENl. The enzyme ligase-1 seals the chain. This is called the ‘long-patch pathway’.
When the initial removal involves lyase action, the endonuclease APE1 recruits DNA polymerase β to replace a single nucleotide. The nick is then sealed by ligase-3. This is called the ‘short-patch pathway’ (Figure 8.12).
Adenine DNA glycosylase recognizes and removes hypoxanthine arising from adenine deamination and alkylated bases such as 3-methyladenine and 7-methylguanine. The methylated adenine is flipped out of the helix into the glycosylase active site and is removed.
Figure 8.12 Long patch and short patch base excision repair
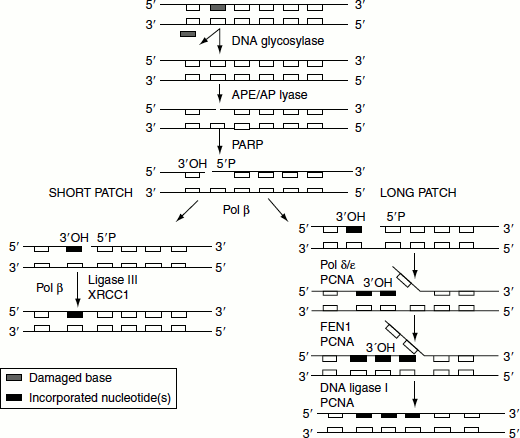
Figure 8.13 Base excision of uracil
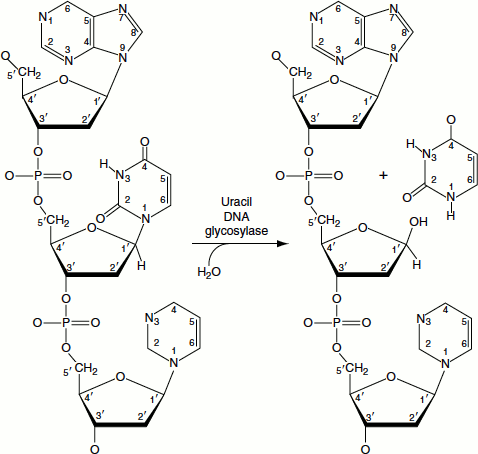
The spontaneous deamination of cytosine results in the formation of uracil in DNA. Such uracil residues are removed by the action of uracil DNA glycosylase (Figure 8.13).
When a base is removed from DNA, the reaction is followed by the excision of the phosphodiester backbone by endonuclease, DNA synthesis by polymerase to fill the gap and ligation by ligase thus restoring the DNA, after correcting its damage.
Nucleotide Excision Repair
DNA damages that cause large distortions in the helical structure are repaired by the nucleotide excision system. In E. coli, the key enzyme is made up of three subunits that are encoded by three genes namely uvrA, uvrB and uvrC. The proteins encoded by these genes uvrA, uvrB and uvrC constitute the ABC excinuclease.
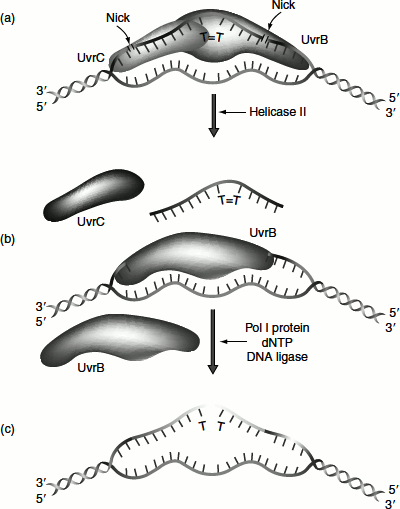
First UvrAB dimer recognizes DNA lesions by an ATP-dependent reaction. UvrA then dissociates and UvrC joins UvrB. The UvrBC complex makes incision on both side of the DNA damage. The incision is made exactly seven nucleotides from the 5′ side of the damaged site and three to four nucleotides away from the 3′ side (Figure 8.14). This incision also requires ATP. UvrD is a helicase that helps to unwind the DNA and helps in the release of the single strands between the cuts. DNA polymerase I (DNA Pol-I) fills the gap that is created by the removal of the damaged strand and ligase seals the gap. In most of the cases, the average length of the replaced DNA is 123 nucleotides (short patch repair). In a few cases, the replacement is nearly 1,500–9,000 nucleotides long (long patch repair).
Figure 8.14 Nucleotide excision repair in prokaryotes
Eukaryotic nucleotide excision repair
The general principles of excision repair in eukaryotic cells are similar to that of bacteria. There are two major pathways of nucleotide excision repair in eukaryotes, which are global genome repair and transcription-coupled repair.
- ‘Global genome repair’ recognizes damage anywhere in the genome. Genes calledXPA to XPG are involved. The XPC protein detects the damage and initiates the repair pathway.
- ‘Transcription-coupled repair’ is responsible for repairing lesions that occur in the transcribed strand of active genes. In this case, the damage is recognized by RNA polymerase II (RNA Pol-II) itself, which stalls when it encounters bulky lesion.
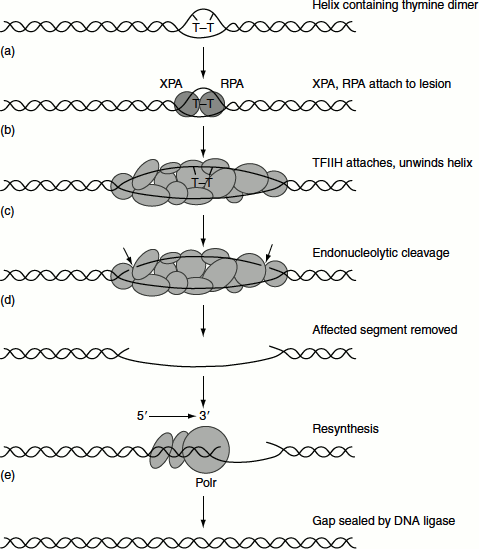
The two pathways eventually merge and use a common set of protein to bring about the repair. The strands of DNA are unwound by approximately 20 bp around the damaged site by the helicase activity of the transcription factor TFIIH, which includes the products of two XP genes, XPB and XPD. XPB helicase is required for promoter melting during transcription, while XPD helicase performs the unwinding function in NER. Cleavages are on either side of the lesion by endonuclease encoded by XPG and XPF genes. About 25–30 nucleotides are excised during NER. The single-stranded stretch including the damaged bases can then replaced by new synthesis and ligated by ligase III and XRCC1 complex (Figure 8.15).
Mutations in the XPD component of TFIIH cause three types of human diseases namely Cockyane’s syndrome, trichothiodystrophy and hereditary non-polyposis colon cancer. XPD is required for the stability of TFIIH complex and the mutations that prevent XPD from stabilizing TFIIH cause trichothiodystrophy. The helicase activity of XPB and XPD is required for repair function. Mutations that affect this helicase activity cause Cockyane’s syndrome.
Figure 8.15 Nucleotide repairexcision repair of eukaryotes
Mismatch Repair
‘Mismatch repair’ is used to repair errors that occur during DNA synthesis. Proofreading during replication is good but not perfect. DNA Pol-III allows the wrong nucleotide to be incorporated about once in every 108 bp synthesized in E. coli. However, the measured mutation rate in bacteria is as low as one mistake per 1010 or 1011 bp. The mismatch repair system is responsible for this final degree of accuracy. Misincorporated nucleotides are recognized and excised and replaced with the correct nucleotides. In contrast to nucleotide excision repair, mismatch repair does not operate on bulky adducts or major distortions of the DNA helix. Most of the mismatches are substitutes within a chemical class, e.g., a C incorporated instead of a T. This causes only a subtle helical distortion in the DNA and the misincorporated nucleotide is a normal component of DNA. The protein ‘mutS’ can distinguish normal base pairs from those resulting from misincorporation.
In E. coli, the parental strand is marked by the methylation of A in a GATC motif, thus the methylation of DNA is used to discriminate parental strands from progeny strands. The ‘dam methylase’ catalyses the transfer of a methyl group to the A of the sequence GATC in duplex DNA. Methylation is delayed for several minutes after replication. In this interval before methylation of the new DNA strand, the mismatch repair system can find mismatches and direct its repair activity to nucleotides on the unmethylated and newly replicated strand. Thus, replication errors are removed preferentially (Figure 8.16).
Figure 8.16 Mismatch repair by mutHLS
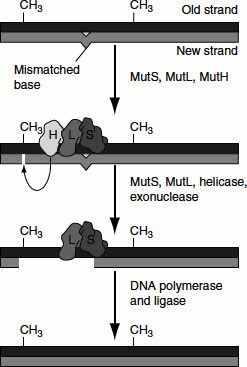
The enzyme complex mutH-mutL-mutS, or mutHLS, catalyses mismatch repair in E. coli. The genes that encode these enzymes are mutH, mutL and mutS. mutS recognizes seven of the eight possible mismatched base pairs (except for C:C) and binds at that site in the duplex DNA. mutH and mutL (with ATP bound) then join the complex and move along the DNA in either direction until they find a hemimethylated GATC motif, which can be as far a few 1,000 bp away. At this point, the nuclease function of mutH is activated in the presence of ATP at a hemimethylated GATC. It cleaves the unmethylated DNA strand, leaving a nick 5′ to the G on the strand containing the unmethylated GATC (i.e., the new DNA strand). The same strand is nicked on the other side of the mismatch.
The segment of single-stranded DNA containing the incorrect nucleotide is excised by UvrD, also known as helicase II and mutU. SSB and exonuclease I are also involved in the excision. As the excision process forms the gap, it is filled in by the concerted action of DNA Pol-III.
Eukaryotic mismatch repair system
Homologous to the E. coli genes, mutL and mutS have been identified in many other species, including mammals. The analysis of mutations that cause one of the most common hereditary cancers, HNPCC, reveals its link to mismatch repair. Some of the genes that, when mutated, cause this disease encode proteins whose amino acid sequences are significantly similar to those of two of the E. coli mismatch repair enzymes. The human genes are called hMLHI (for human mutL homologue 1), hMSHI and hMSH2 (for human mutS homologue 1 and 2 respectively).
Recombination Repair
In order to understand the mechanism of recombination repair, it is necessary to know the effect of thymine dimers on DNA replication. When polymerases reach a thymine dimmer, the replication fork is temporarily stalled. A thymine dimmer is still capable of hydrogen bonding with two adenine bases, because dimerization does not alter the hydrogen bonding groups of thymine. However, such dimerization introduces a distortion in the DNA helix and when polymerase adds adenine to the growing chain, it reacts to the distortion as if a mismatched base had been added and thereby removes the added adenine by its proofreading activity. This cycle of adenine addition and removal against the thymine dimer continues and as the result the polymerase stalls at this site. This stalling can be overcome by two ways called ‘post-dimer initiation’ and ‘trans-dimer synthesis’ and these events are responsible for ‘recombination repair’ and ‘SOS repair’.
In recombination repair, the thymine dimers are bypassed and chain growth is initiated beyond the block. In such post-dimmer initiation, the replicated daughter strands are different in nature. One has the parental strand containing the damaged adduct and hence this daughter will have large gaps, one for each unexcised dimer. The other daughter strand has the undamaged parental strand and this daughter DNA will be copied normally. The gap opposite the damaged site in the first duplex is filled by the homologous single strand of DNA from the normal duplex and this process is called ‘sister-strand exchange’.
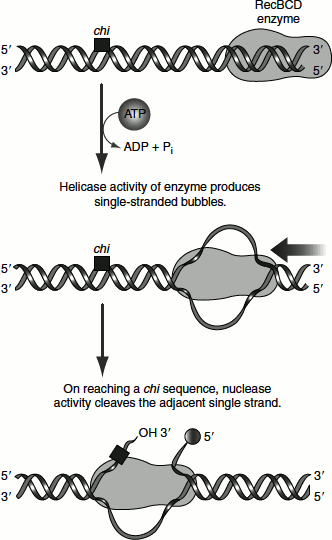
In E. coli, the recB, recC and recD genes encode the RecBCD enzyme, which has helicase and nuclease activities. The RecBCD binds to linear DNA at a free (broken) end and moves inward along the double helix, unwinding and degrading the DNA in a reaction coupled to ATP hydrolysis. The enzyme nears a sequence called chi (5′) GCTGGTGG (Figure 8.17). From this point, the degradation of the strand with a 3′ terminus is greatly reduced but degradation of the 5′ terminal strand is increased. This process creates single-stranded DNA with a 3′ end, which is used during subsequent steps in recombination.
Figure 8.17 Recombination repair assisted by the chi sequence and the RecBCD enzyme
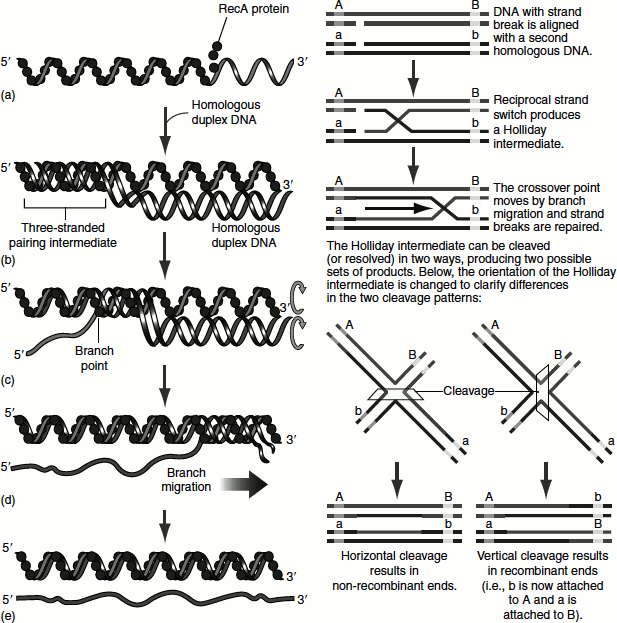
The RecA monomers assemble cooperatively on DNA and form a helical filament. The RecF, RecO and RecR proteins regulate the assembly of RecA filaments. As the duplex DNA is incorporated within the RecA filament and aligned with the bound single-stranded DNA, one strand of the duplex switches pairing partners. The the duplex switches pairing partners. The continued strand exchange requires an ordered rotation of the two aligned DNAs. This results in a spooling action that shifts the branch point along the helix. ATP hydrolysis by RecA provides the necessary energy Figure 8.18 (A). The resulting product is called a ‘Holliday intermediate’ Figure 8.18 (B). Once a Holliday intermediate is formed, enzymes such as topoisomerases, the RuvAB branch migration protein, resolvase, nuclease, polymerase and ligase complete the recombination reaction. The RuvC protein of E. coli cleaves Holliday intermediates to generate full-length and unbranched recombined and repaired chromosome products.
Figure 8.18 (A) Recombination repair (B) Homologous recombination
Recombination repair in eukaryotes
The RAD 52 group of genes is required for recombination repair in eukaryotes. The MRX (yeast) or MRN (mammals) complex is required to form single-stranded region at each DNA end. The RecA homologue Rad51 forms a nucleoprotein filament on the single-stranded regions, assisted by Rad 52 and Rad 55/57. Rad 54 and Rdh54/Rdh54B are involved in homologue search and strand invasion. Following repair synthesis, the resulting structure is resolved.
SOS Repair/Error-Prone Repair
‘SOS repair’ is a bypass system that allows DNA chain growth across damaged segments. It is an error-prone process. Even though the DNA strands are repaired, they contain incorrect bases. The principle involved is that survival with mutations is better than no survival at all. For this reason, SOS repair is also known as ‘error-prone translesion DNA synthesis’. In bacteria, this repair takes place in response to extensive DNA damage known as SOS response.
SOS response
Agents that damage the DNA, such as UV radiation, alkylating agents and cross-linking agents, induce a complex system of cellular changes in E. coli known as the SOS response. E. coli cells so treated stop dividing and increase their capacity to repair the damaged DNA. The damaged DNA cause RecA to trigger the SOS response and results in the autocleavage of a protein called LexA protein between its Asp 84 and Gly 85. RecA is activated on binding a single-stranded DNA. LexA functions as a repressor of 43 genes that participate in DNA repair and the control of cell division including recA, lexA, uvrA and uvrB. The LexA repressible genes are preceded by the ‘SOS box’, a palindromic homologous 20 nucleotide sequence. LexA specifically bind to the SOS boxes of recA and lexA.
During normal growth, LexA repress the expression of SOS genes, including the lexA gene by binding to their SOS boxes and prevents RNA polymerase binding and transcribing these genes.
Figure 8.19 SOS response
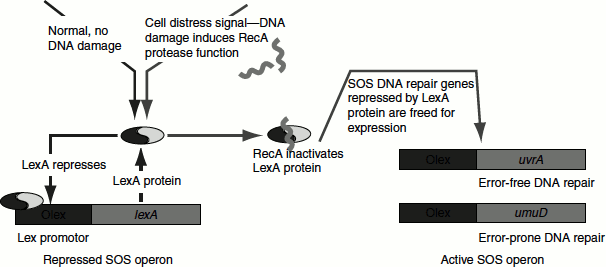
When DNA damaged is extensive and produces single-strand gaps, they bind to RecA and stimulate LexA cleavage. As the result, the genes repressed by LexA are released from repression and induce the synthesis of SOS proteins. When DNA lesions have been eliminated, RecA ceases the stimulation and stops the autoprotolysis of LexA. The newly synthesized LexA can then function as a repressor and the cell thus returns to normality (Figure 8.19).
The E. coli Pol-III holoenzyme is unable to replicate through a variety of lesions such as AP sites and thymine dimers. On encountering such lesions, the replisome stalls and Pol-III core enzyme released. This process is called ‘replicative fork collapse’. Cells have two general modes for restoring collapsed replication fork, which are recombination repair and SOS repair.
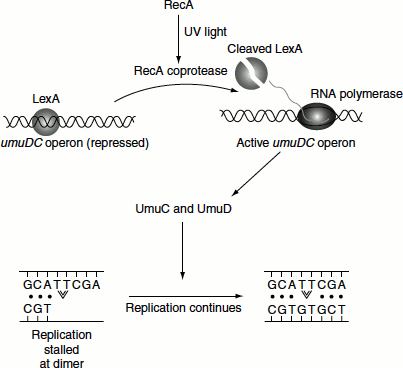
In SOS repair, the Pol-III core lost from the collapsed replication fork is replaced by by-pass DNA polymerases namely DNA Pol-IV and DNA Pol-V. Both of these enzymes are Y-family DNA polymerases and lack the 3′ → 5′ proofreading activity. Thus, they replicate the undamaged DNA with poor fidelity and low processivity. Thus, SOS repair is also known as error-prone repair. DNA polymerase is a 336-residue product of dinB gene. DNA Pol-V is a heterotrimeric product of UmuD and UmuC genes and Umu D′2C (Umu means UV-assisted mutagenesis). UmuD′ is produced by the RecA-assisted cleavage of UmuD (Figure 8.20).
Translesion synthesis (TLS) by Pol-V, which is also called Pol-V mutasome, tends to incorporate G about half as often as A, opposite to thymine dimers and AP sites. This process is highly mutagenic. After seven nucleotides are synthesized, Pol-V mutasome is replaced by Pol-III holoenzyme, which normally continues the replication process. Pol-II also is induced by SOS response and participates in TLS. The role of Pol-II is to mediate an error free TLS and only if this process fails, it is replaced by Pol-V to carry out the error-prone TLS.
Figure 8.20 RecA relieves the repression of SOS box by LexA cleavage
SOS repair is a mutagenic process and is, therefore, a process of last resort that is initiated when the DNA is repaired by other repair systems. Indeed most mutations in E. coli arise from the action of SOS repair system. Under conditions of environmental stress, the SOS system functions to increase the rate of mutation to increase the rate at which E. coli adapts to new conditions.
Eukaryotic polymerase η, ι, κ and ζ are implicated in TLS.