The term restriction endonuclease was coined by Lederberg and Meselson in 1964 to describe the nuclease enzyme that destroys or restricts any foreign DNA entering a bacterial cell. These restriction endonucleases are widely used in rDNA technology. They specifically bind to double-stranded DNA and cleave it at specific sites known as recognition sequence or restriction sites. They recognize specific sequences that are 4–6 bp in length and show a two-fold dyad symmetry. The DNA fragments produced by the action of restriction endonuclease help in the joining of DNA fragments to form new rDNA.
Types of Restriction Endonucleases
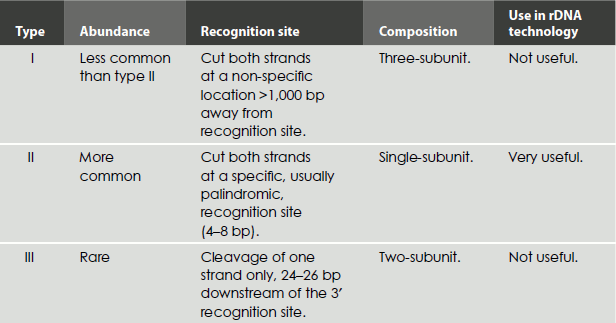
Restriction endonucleases are of three types, namely types I, II and III (Table 9.1). Their grouping is based on the types of sequences recognized, the nature of the cut made in the DNA and the enzyme structure. Type I and type III restriction endonucleases are not useful for gene cloning, because they cleave DNA at sites other than the recognition sites and thus cause random cleavage patterns. In contrast, type II endonucleases are widely used for mapping and reconstructing DNA in vitro because they recognize specific sites and cleave just at these sites.
Table 9.1 Types of restriction endonuclease
Type I Enzymes : These enzymes attach to DNA molecule and migrate to a distance of about 1,000–5,000 nucleotides and cleave the DNA strand at a random site, creating a gap of about 75 nucleotides in length. Type I enzymes also methylate DNA, i.e., they function both as endonuclease and as methylase. They consist of three subunits—one subunit is responsible for restriction, one subunit for methylation and the third subunit for DNA binding. Examples of type I restriction enzymes are EcoB and EcoK.
Type II Enzymes : These enzymes are used for gene manipulation studies. They recognize specific target sequence and cleave the double-stranded DNA molecule within or near the recognition sequence. Their action results in DNA fragments of defined length and sequence. They require Mg2+ ions for their activity. Example of type II enzymes are EcoRI, EcoRII, etc.
Type III Enzymes : These enzymes consist of three subunits—one specifies site recognition, one specifies methylation and the other specifies cleavage. These require ATP as the source of energy and Mg2 ions as cofactor. They cleave DNA at specific non-palindromic sequences. Example of type III enzymes are HpaI, MboII, etc.
Nomenclature of Restriction Endonuclease
The restriction enzymes are named based on the following principles:
- Restriction endonucleases are named for the organism in which they were discovered. The name of the organism is identified by the first letter of genus name and first two letters of species name to form a three-letter abbreviation, which is italicized.For E. coli = EcoFor H. influenzae = Hind
- A strain or type of the organism in which the restriction enzyme is identified is also written along with the name.
- For example, EcoK for E. coli strain K, Hind for H. influenzae strain Rd.
- The restriction systems genetically specified by plasmid of the organism is also indicated. For example, EcoRI, EcoPI.
- When a strain has several restriction and modification systems, they are identified by Roman numerals. For example, Hind I, Hind II and Hind III. These should not be confused with the Roman numerals used for specifying the types.
Restriction enzymes and their recognition sequences
| Name | Source microorganism | Recognition sequence |
|---|---|---|
| Bam HI | Bacillus amyloliquefaciens | G↓GATCC |
| ECo RI | Eschericia coli RY13 | G↓AATTC |
| Hind III | Haemophilus influenzaeRd | A↓AGCTT |
| Not I | Nocardia otitidiscaviarum | GC↓GGCCGC |
| Pst I | Providencia stuartii | CTGCA↓G |
| Sma I | Serratia marcescens | CCC↓GGG |
Type II Restriction Endonucleases
Restriction enzymes can cut to produce:
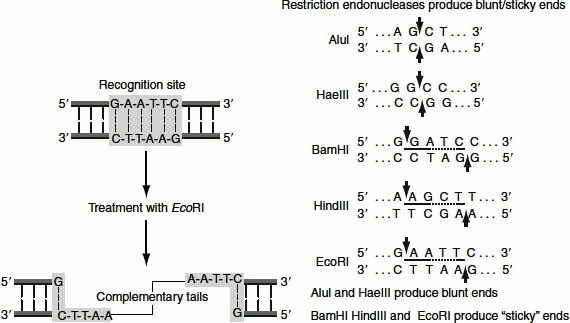
- ‘Blunt ends or flush ends’, i.e., they cleave both strands of DNA at same base pairs, in the centre of the recognition sequence (Figure 9.5).For example, Hae III (Haemophilus aegypticus, order of the enzyme III) recognizes a four-nucleotide-long palindromic sequence and cuts symmetrically both DNA strands forming blunt ends. It recognizes the sequence
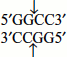 The cut is made between the adjacent G and C.
The cut is made between the adjacent G and C. - ‘Sticky ends or cohesive ends’, i.e., they make staggered cuts; for example, EcoRI recognizes the sequence (Figure 9.5).
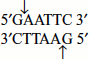
The cut is made between G and A residues of each strand and produces two single-stranded complementary cut ends that are asymmetrical having 5′ overhangs of four nucleotides.
Figure 9.5 Restriction endonucleases producing blunt and sticky end
Isoschizomers
In general, different restriction enzymes recognize different sequences. However, a few of them recognize the same restriction sites. Such restriction endonucleases that are isolated from two different sources but possess similar recognition and cleavage sites are called ‘isochizomers’.
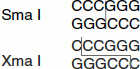
Isoenzymes
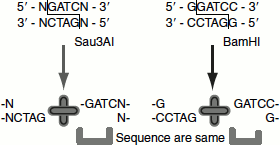
If two different restriction endonucleases produce same cohesive ends, then the two enzymes referred as isoenzymes.
For example, Bam HI and Sau3AI create the fragments with cohesive end of ‘GATCC’ by recognizing the sequence of GGATCC and NGATCN′, respectively (Figure 9.6).
Figure 9.6 Isoenzymes