If DNA is denatured and later allowed to renature, the two separated single strands of DNA will zipper back to reform the double-stranded DNA molecule. This ability of the complementary sequences to anneal or to hybridize one another is called ‘nucleic acid hybridization’. This technique helps in determining the gene structure and in identifying molecules that contain the same nucleotide sequence. Thus, from a complex mixture of nucleic acid molecules, hybridization techniques help in the separation of complementary sequence.
Hybridization is normally performed using one labelled sequence called the ‘probe’. Probe is a short oligonucleotide that is complimentary to the target DNA sequence.
For the identification of hybridized nucleic acid duplexes, the labelling of probe is necessary. There are various ways of labelling nucleic acids, which are:
- Nick translation,
- End labelling or end filling,
- Non-radioactive labelling
- Biotin labelling
- Digoxigenin labelling,
- Fluorescein labelling and
- Enzyme labelling.
Nick translation
E. coli DNA Pol-I adds nucleotides to the 3’-OH terminus that is created when one strand of the double-stranded DNA molecule is nicked. In addition, polymerase I can remove nucleotides from the 5’-side of the nick. The simultaneous removal of nucleotides from the 5’-end of the nick and the addition of nucleotides to the 3’-end of the nick result in the movement of the nick (nick translation) along the DNA. By replacing the pre-existing nucleotides with 32P-nucleotides, the 32P-labelled DNA can be obtained (Figure 9.24).
End labelling or end filling
This is a gentler method that rarely causes a breakage of the DNA (nick translation can cause the breakage of DNA under some circumstances). This method can be used to label DNA molecules that have sticky ends. The Kelenow fragment of DNA Pol-I is used. If the reaction is carried out in the presence of labelled nucleotides, then the DNA is labelled (Figure 9.25).
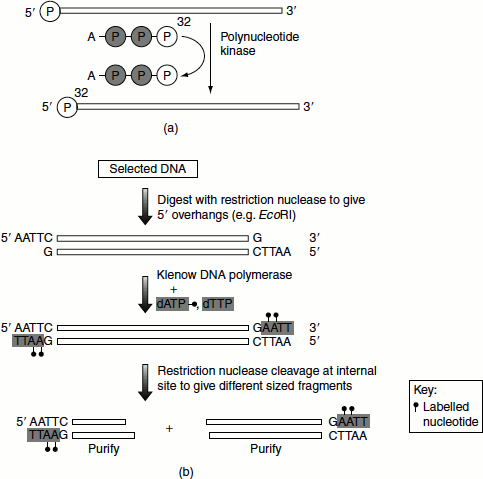
Kinase end labelling
The 5’ terminal phosphate is replaced by 32P-labelled γ-phosphate of [γ-32P] ATP (Figure 9.25(a)).
Fill-in end-Labelling
The DNA of interest is cleaved with a suitable restriction endonuclease to generate sticky ends. These sticky ends act as primer for Klenow DNA polymerase and incorporate labelled nucleotides (Figure 9.25(b)).
Figure 9.24 Nick translation
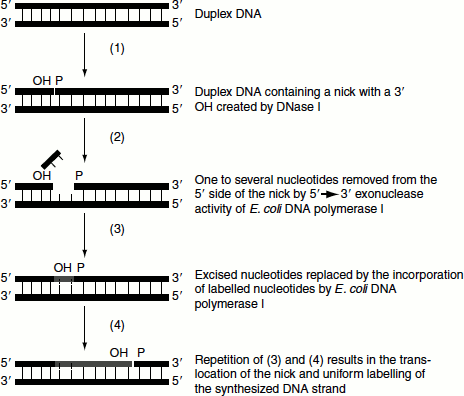
Figure 9.25(a) Kinase end labelling of oligonucleotides and (b) Fill-in end-labelling
Non-radioactive labelling
Biotin labelling
dUTP molecules are modified by the reaction with biotin. By nick translation, it is possible to introduce 10–15 biotin dUTP per 100 nucleotides. The biotinylated probe is then be detected by treatment with avidin coupled to a fluorescent marker. Biotinylated adenosine and cytosine triphosphates can also be used instead of dUTP.
Fluorescein labelling
Fluorescein nucleotides such as fluorescein dUTP/UTP/ddUTP are incorporated into oligonucleotides by nick translation. Detection is done by an anti-fluorescein antibody, which is coupled to an enzyme.
Enzyme labels
Enzymes such as peroxidase and alkaline phosphatase can be coupled directly to DNA probes in the presence of glutaraldehyde. The enzyme-labelled probe is hybridized with a target DNA and visualized by adding its chromogenic substrate.
Advantages of non-radioactive labels
- They are safe than radio-labels.
- They have long shelf life.
- There is no radioactive waste disposal problems.
- They can be used in concentration that shortens the hybridization time.