Genomic Library
It is a collection of clones containing every single gene present in an organism. For the construction of a genomic library, the entire genomic DNA is isolated from host cells/tissues, purified and broken randomly into fragments of appropriate size for cloning into suitable vector. DNA can be fragmented by physical shearing or by the action of restriction enzymes. These experiments with randomly cloned fragments are known as ‘shot gun cloning experiment’.
The various steps involved in creating a genomic library can be discussed as follows:
- The chromosomal DNA of the organisms is isolated.
- It is then treated with a known restriction enzyme.
- The fragments are cloned using appropriate cloning vector.
- For identifying recombinants, a marker gene is inserted while multiplying in vector.
- The rDNA is then hybridized by two major techniques, namely colony hybridization and plaque hybridization.
Colony hybridization
The cloned colonies are transferred from culture plate onto a nitrocellulose filter paper. The filter with the colony replicas is treated with NaOH to lyse the host cells and to denature the DNA. The filter is then baked to fix the DNA. It is then treated with a radio-labelled probe that is complementary to the DNA of interest. The filter is then washed to remove the unbound excess probe. It is then autoradiographed. This indicates the host colonies that carry the desired gene. The colony is taken out from the master culture plate and then mass cultured.
Plaque hybridization
This technique is used when a phage particle is carrying the gene of interest. In this case, a culture of bacterium is infected with a mixture of chimeric phage particles. A large number of plaques develop overnight. These plaques are then treated like the colonies in colony hybridization, thus identifying and isolating the chimeric phage particle carrying the gene of interest.
As the genomic DNA of eukaryotes contains more introns, regulatory regions and repetitive sequences, the establishment of genomic library of eukaryotes is not meaningful. Hence, the cDNA library is generally established for eukaryotes.
cDNA library
For higher organisms such as plants and animals, a gene library will contain so many different clones, so that the identification of a desired clone is difficult. Here, a cDNA library is useful. Since only a few genes are expressed in any cell and only those genes that are expressed are transcribed into an mRNA, if the mRNA is used as the starting material, the resulting clones will comprise only a selection of the total number of genes in the cell. The mRNAs cannot be directly cloned because they are unstable and hence they are converted into their complementary DNAs (cDNAs). The library made from comple-mentary or copy DNA is called ‘cDNA library’. The cDNA library can be made from mRNA because they are highly processed, intron-free and have only coding sequences.
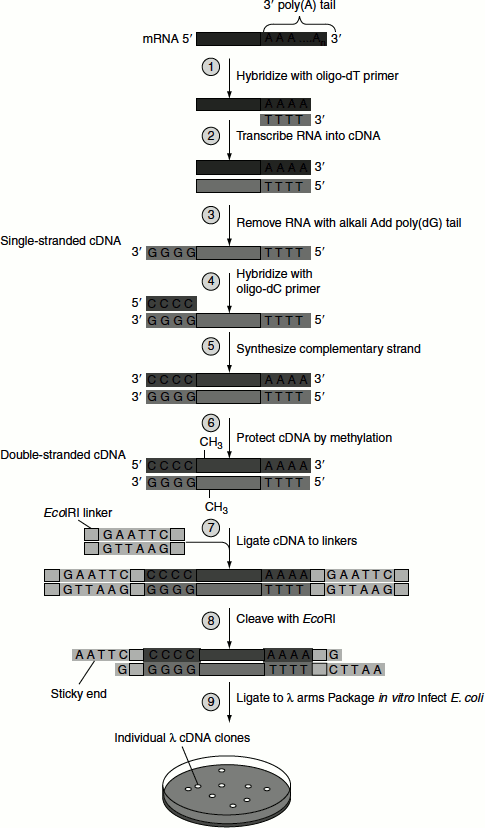
Steps involved in establishing cDNA library are (Figure 9.30):
- Isolation of mRNA : The majority of mRNA sequence in eukaryotes contains a long polyadenylated tract at their 3’-end. Therefore, mRNA binds to an oligo-dT cellulose affinity column or poly-U sepharose column from which it can be eluted.
- Reverse transcription : Reverse transcriptase is required for the synthesis of DNA copy of an mRNA. The mRNAs are treated with oligo-dT primer, reverse transcriptase enzyme and dNTPs. The oligo-dT primer binds to polyadenylated tail and provides free 3’-OH for reverse transcription. Reverse transcriptase adds complementary dNTPS one by one to the free 3’-OH group of the primer and thereby results in the formation of RNA-DNA hybrid. Cellular DNA and total RNA inhibit reverse transcriptase; hence, it is necessary that the mRNA must be in pure form before cloning.
- Oligo-dC tailing : The RNA–DNA hybrid is treated with the enzyme terminal transferase and dCTP. This enzyme adds dCTP one by one to the 3’-OH group of RNA and DNA strands. As the result, a short oligo-dC tail is produced at the 3’-end of both the strands. The cDNA becomes now curved, forming a hairpin loop.
- Alkali hydrolysis : On treatment with alkaline sucrose solution, the mRNA–cDNA strands are separated into single strands.
- Addition of oligo-dG primer : Oligo-dG primer is then added to the reaction mixture and the temperature is maintained at 55°C. This favours the binding of dG primer to oligo-dC tails formed on cDNA. Now, cDNA acts as a template for the synthesis of double-stranded cDNA in the presence of DNA Pol-I.
- Cloning of cDNA : The blunt-ended cDNA is treated with linkers/adaptors and subjected to restriction enzyme action creating sticky ends that are ligated to appropriate vector and transferred into a bacterium. Each bacterial cell possesses a single-stranded cDNA clone and hence the collection of all recombinant bacteria is called cDNA library.
- Screening of cDNA clones : The cDNA clones can then be selected by colony hybridization.
Figure 9.30 cDNA library