Euchromatin and heterochromatin can be localized during the interphase in the nucleus. Heterochromatin is highly inert and condensed, transcriptionally repressed, replicates during the late S-phase and is found in the periphery of the nucleus. Centromeric heterochromatin predominantly consists of satellite DNAs. When a gene is transferred to a position adjacent to the heterochromatin, it may also become inactive, i.e., it too becomes heterochromatic. Such inactivation is the result of an ‘epigenetic effect’. It differs between individual cells in an animal. This results in the phenomenon of ‘position effect variegation’, where genetically identical cells have different phenotypes; for example, the position effect variegation in the fly eye. A few regions in the eye are colourless, whereas a few others are red in colour. This is because the white gene required for developing the red pigment was inactivated by heterochromatin in some cells, while remained active in others.
Inactivation spreads from heterochromatin into adjacent region for varying distance, which may extend from a nearby gene to distant gene. This inactivation occurs during the embryonic state and thereafter is stably inherited by all progeny cells.
The formation of heterochromatin is a two-stage process that includes:
- A nucleation event occurs by the binding of a protein that recognizes the heterochromatic sequence.
- Propagation of the nucleated structure throughout the chromatin fibre.
However, such spreading of heterochromatic regions is prevented by the presence of an activated promoter of the nearby gene and by the presence of insulators.
Telomeric Silencing
Genes translocated to a telomeric location shows variable loss of activity. This results from the spreading effect that starts from the telomeres. The binding of a protein called Rapl to the telomeric sequence triggers the nucleation event, which recruits the heterochromatin proteins thereby inactivating the gene sequences.
The inactivation of chromatin occurs by the addition of proteins to the nucleosomal fibre that results in chromatin condensation thus making it inaccessible to the transcription machinery. The proteins added to the heterochromatic DNA may also directly block the regulatory sites or sometimes even can directly inhibit transcription. The molecular mechanisms needed for the formation of heterochromatin has been identified in Drosophila mutants. They include the following genes.
- Su(var)—The gene products of this gene suppress the position effect variegation.
- E(var)—The gene products of this gene enhance the position effect variegation.
The genes were so named for the behaviour of the mutant loci. The Su(var) mutations lie in the genes whose products are needed for the formation of heterochromatin. They include enzymes that act on chromatin-like histone deacetylase. The E(var) mutations lie in the genes whose products are required for activating gene expression. They include the members of SWI/SNF complex. HP1 (heterochromatic protein 1) is one of the most important Su(var) proteins.
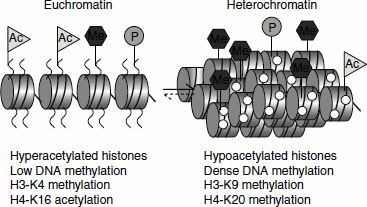
The state of histone methylation is an important factor that determines whether a DNA sequence exists as heterochromatin or euchromatin state. The methylation of histone H3 lysine 9 favours heterochromatin formation, while histone H3 lysine methylation is required for euchromatin formation (Figure 10.1). A trimethyl H3K4 demethylase known as Lid2 in Schizo-saccharomyces pombe interacts with H3K9 methyltransferase resulting in H3K4 hypomethylation and heterochromatin formation. The link between H3K4 demethylation and H3K9 methylation suggests that the two reactions act in a co-ordinated manner to control the hetero-chromatic and euchromatic state of a specific region.
Figure 10.1 DNA methylation and histone modifications
Heterochromatin formation at telomeres depends on a set of genes known as ‘silent information regulators’ (SIR genes). The mutations of SIR genes relive the inactivation of the genes that are integrated near telomeric heterochromatin. Rap1 has the crucial role in the formation of heterochromatin. It recruits Sir4, which in turn recruits Sir3 and HDAC. Sir2, Sir3 and Sir4 then interact directly with the histones H3 and H4. This complex can polymerize further and gradually spreads along the chromatin. This process inactivates the DNA region because coating with Sir3/Sir4 complex has an inhibitory effect or because Sir2-dependent deacetylation that represses the transcription.
Silencing complexes repress chromatin activity because they condense chromatin, so that regulatory proteins cannot find their target sequences. Further, the presence of these proteins hinders the binding of transcription factors and RNA polymerase. They also block chromatin remodelling. The gene activation and the repressing effects of chromatin compete with each other and thus the activation of a promoter of an adjacent gene inhibits the spread of silencing complex.
In human cells, the centromeric specific protein CENP-B is required to initiate histone modification (H3 deacetylation of K9 and K14, followed by methylation of K9) that triggers the binding of HP1 and ultimately leads to the formation of heterochromatin in that region.