DNA methylation occurs at specific sites. In bacteria, the DNA methylation is used for identifying bacterial restriction-methylation system that is involved in phage defence and is also used for distinguishing replicated and non-replicated DNA. In eukaryotes, the DNA methylation is connected with the control of transcription. The methylation of a control region is usually accompanied by gene inactivation. Methylation in eukaryotes usually occurs at the CpG islands in the 5′ regions of some genes. These are the CG dinucleotide-rich regions of the gene. About 2–7 per cent of animal cell DNA is methylated. Methylation occurs at the 5th position of cytosine producing 5-methylcytosine. Most methyl groups are found in CG dinucleotides in CpG islands, where the C residues on both strands of this short palindromic sequence are methylated. Such a site is considered fully methylated. Upon replicating this site, each daughter duplex will have one methylated strand and one unmethylated strand. Such site is called hemimethylated. If methylation of the unmethylated strand occurs, then the hemimethylated site also becomes fully methylated.
If replication occurs first, then the hemimethylated condition will be perpetuated on one daughter duplex and the other duplex will become unmethylated.
The state of methylation is controlled by DNA methyltransferases or methylases or DNMTs, which adds methyl groups to the 5th position of cytosine (Figure 10.2). There are two types of methyltransferases, namely:
- De novo methyltransferase: It modifies DNA at a new position and acts only on unmethylated DNA. There are two de novo methyltransferases, namely DNMT3A and DNMT3B in mouse; these have different target sites.
- Maintenance methyltransferase: It acts constitutively only on hemimethylated DNA and converts them to fully methylated sites.
A protein called UHRF1 is important for the maintenance of methylation and associates with DNMT1. This protein is able to recognize CpG dinucleotides and preferentially binds to hemimethylated DNA. UHRF1 increases the efficacy of DNMT1 for maintenance methylation at hemimethylated sites. UHRF1 also interacts with methylated histone H3. This shows that the maintenance of DNA methylation is connected with the stabilization of heterochromatin structure.
Promoters are the most common sites of methylation. They are methylated when gene is inactive and unmethylated when gene is active. DNA methylation plays a role in controlling gene expression. The transcriptional silencing of selected genes caused by DNA methylation plays a crucial role in the development and progression of human cancers.
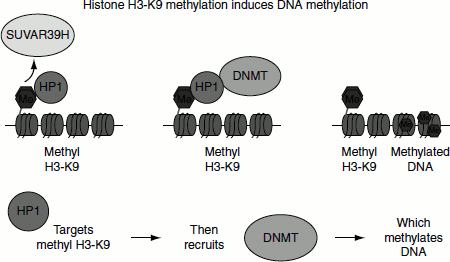
Figure 10.2 SUVAR39H is a methyltransferase that specifically methylates the lysine 9 of histone H3. Such a methylation creates a binding site for the heterochromatin protein HP1 that recruits a DNA methyltransferase, capable to methylate the CpG in DNA (Me = methyl; methyl H3-K9 = methyl on lysine 9 of histone H3; HP1 = heterochromatin protein 1; DNMT = DNA methyltransferase)
Satellite DNAs are also methylated. The mutations of DNMT3B prevent the methylation of satellite DNA and this causes centromere instability; for example, a disease called ‘ICF—immunodeficiency centromere instability facial anomalies’ is caused by such mutations.
‘Rett syndrome’ is another disease that emphasizes the importance of DNA methylations. It is caused by the mutations of the gene for the protein MeCP2 that binds methylated CpG sequences. The disease is characterized by autism-like symptoms that are the result of a failure of normal gene silencing in the brain.
There are several ways to generate a demethylated site. These include:
- Loss of methylation at a site due to the incomplete fidelity of DNMT1 during maintenance methylation.
- Blocking the maintenance methylase from acting on the site when it is replicated. After the second round of replication, the daughter DNA will be unmethylated.
- Actively demethylating the site, i.e., removing methyl groups from methylated cytosine from the DNA and the excised region is then filled by repair system. Then, enzyme cytidine deaminase may be involved, which introduces a mismatch that is then corrected.
Active demethylation can occur to the paternal genome soon after fertilization. DNMT3A and DNMT3B participate in active DNA demethylation. They may possess deaminase activity and are involved in gene demethylation. The enzyme mediates the oxidative deamination of cytosine and converts it to 5-methylcytosine (thymine), the resulting guanine-thymine G-T mismatch is repaired by base excision repair thus leading to the demethylation of previously methylated CpG site.