This is a molecular biological technique used for amplifying specific DNA sequences. The target DNA sequence can be amplified billions of times. A PCR reaction requires target DNA, free nucleotides, DNA primers and the enzyme Taq polymerase. This enzymatically mediated in vitro amplification of specific target sequence is achieved by simultaneous extension of complementary strands of DNA (Figure 12.12). This is carried out by repeated cycles (up to 35 cycles) of heating (95°C), cooling (37°C–65°C) and extension at 72°C using thermostable DNA polymerase. The PCR reaction is carried out in a plastic tube which is inserted in a regulatable heating block called the thermocycler. The PCR machine can be programmed to heating and cooling as per the required order. The target DNA sequence thus gets exponentially amplified.
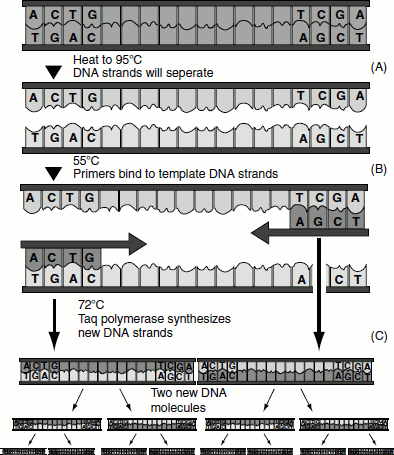
Two primers (short oligonucleotide sequence) called forward and reverse primers, both synthesized in the 5’ → 3’direction are designed in such a way that they are complementary to discrete sequences on the target DNA. By heating, the duplex target DNA is separated. Upon cooling, the primers anneal to the target sequence. In presence of MgCl2, buffer and dNTPs, the bound primers are extended. The products of this PCR reaction is described as first cycle primer extension.
During the second cycle, the double-stranded products of the first cycle are heated and separated into single strands. Each of the single strands act as templates and binds with the primers. The annealed primers are extended by the thermostable polymerase as before. The same process can be continued for third cycle and so on till the desired DNA amplification is reached.
The PCR Reaction Components
Oligonucleotide primers (also called amplimers) dNTPs, thermostable DNA polymerase, reaction buffer and MgCl2.
Figure 12.12 The PCR reaction
Thermostable polymerase
These polymerases are obtained from organisms that thrive in hot springs and have an optimum activity at 72°C. The most commonly used thermostable polymerase is Taq polymerase. It is isolated from the organism Thermus aquaticus. Other thermostable polymerases include Vent polymerase, which is isolated from Thermococcus litoralis, and Tth polymerase, which is isolated from Thermus thermophilus. Usually, about one unit of the enzyme is used per reaction. This can amplify up to 40 cycles.
Target DNA
The sequence of the target DNA can also affect the PCR amplification. If the DNA is GC-rich, then there will be difficulties in separating the DNA strands. In such cases, the addition of formamide or dimethyl sulfoxide (DMSO) can resolve the problem. The concentration of the target DNA also influences the degree of amplification; hence, it is necessary to dilute the target DNA appropriately.
Primers
Primers are chemically synthesized oligonucleotides. The optimum size of a primer is between 18 and 25 nucleotides. The primers are carefully designed, so that there are no inter- and intramolecular complementarities, as such complementarities can lead to primer dimmers which will inhibit the PCR reaction. The primers are also designed in such a way that they do not have a high GC content.
Magnesium chloride
The concentration of MgCl2 is very important for a PCR reaction. MgCl2 is required for DNA polymerase activity. It influences the binding of the primers to the template DNA.
PCR Thermocycling
PCR amplification is achieved by repeated cycles of heating (denaturation), cooling (annealing) and extension. A typical cycle for the amplification of a DNA fragment of 500 bp would be 95°C (denaturation) for 60 s, 50°C (annealing) for 60 s. Larger amplification products need longer denaturation and extension periods. The sequence of primers and ionic conditions will determine the annealing temperature best suited for the cycle.
Types of PCR
There are different types of PCR. Some of the common types of PCR are:
- real-time PCR,
- reverse transcription PCR (RT-PCR),
- multiplex PCR,
- nested PCR,
- quantitative PCR,
- in situ PCR,
- colony PCR and
- standard PCR.
Real-time PCR
This is so called because it allows the scientist to actually view the increase in the amount of DNA as it is being amplified (Figure 12.13). This type of PCR combines the PCR amplification and detection into a single step, thus eliminating the need to detect products using gel electrophoresis and enabling the PCR method to be truly quantitative. This type of PCR is also called quantitative real-time PCR (Q-PCR/qPCR/qrt-PCR) or kinetic PCR (KPCR). There are several types of real-time PCR. The PCR products in real-time PCR can be detected by two common methods. They are:
- Using non-specific fluorescent dyes that intercalate with double-stranded DNA.
- Using sequence-specific probes consisting of oligonucleotides that are labelled with a fluorescent reporter. The probe hybridizes with the target DNA and permits its detection.
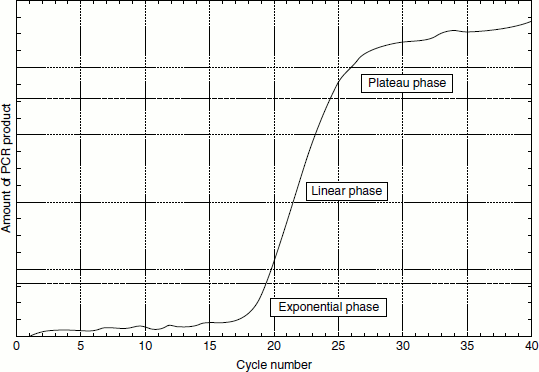
Figure 12.13 Phases of PCR amplification
Real-time PCR has many applications; these include:
- the precise measurement of changes in gene expression,
- overall gene expression quantitation,
- the detection of single nucleotide polymorphisms (SNPs) and
- post-PCR detection for allelic discrimination assays.
The real-time PCR allows the investigator to monitor fluorescence produced by a reporter dye during each cycle. The amount of fluorescence is proportional to the amount of amplicons generated. This process allows for detection of all phases of amplification: exponential, linear and plateau.
Reverse transcription PCR
This is a powerful tool for the detection and quantification of mRNA expression. This technique is so sensitive that even quantitation of RNA from a single cell is possible. RT-PCR involves two steps; they are:
- The reverse transcriptase reaction and
- PCR amplification.
In this process, RNA is first reverse transcribed into complementary DNA (cDNA). The resulting cDNA is used as templates for subsequent PCR amplification using specified primers. RT-PCR can be carried out as a one-step reaction or a two-step reaction (Figure 12.14).
Figure 12.14 RT-PCR
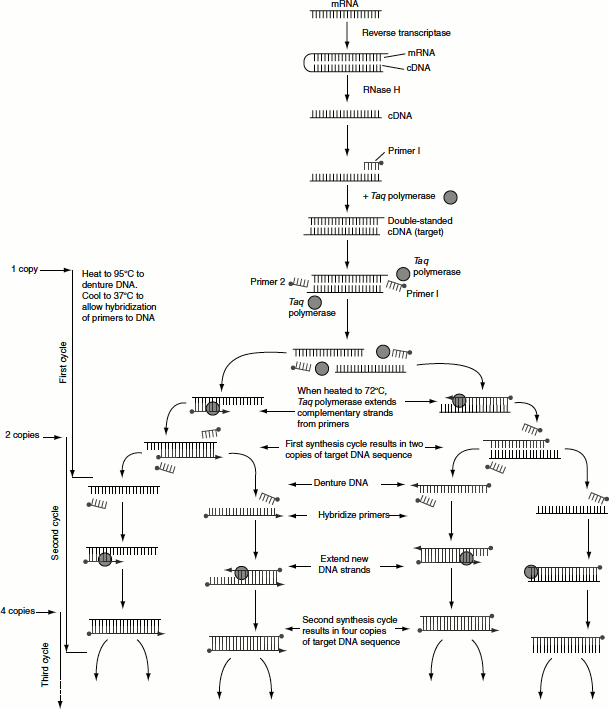
In one-step reaction, all reaction components are mixed in one tube prior to the initiation reaction. The disadvantage with one-step reaction is that the resulting cDNA cannot be used for detecting multiple messages from a single RNA sample as in two-step RT-PCR. RT-PCR-based assays are the most common method for characterizing gene expression patterns and for comparing mRNA levels in different sample population.
Multiplex PCR
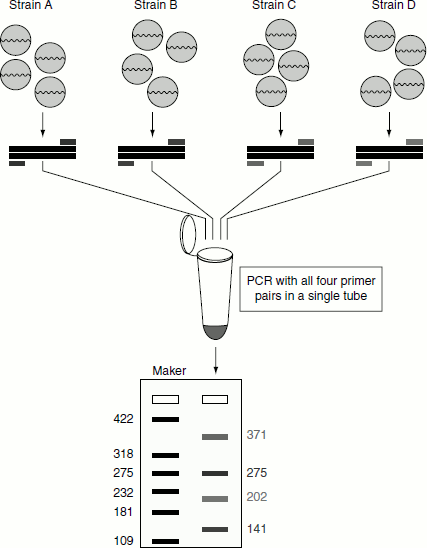
This type of PCR enables the amplification of multiple targets in a single PCR experiment. In this type of assay, more than one target sequence can be amplified by using multiple primer pairs in the PCR reaction mixture (Figure 12.15). Multiplexing reactions are of two types, namely:
- Single template PCR reaction: Here, a single template (which can be genomic DNA) is used along with several pairs of primers that amplify specific regions of the template.
- Multi template PCR reaction: This type of reaction uses multiple templates and several primer sets in the same reaction tube. However, the presence of multiple primers may lead to cross hybridization with each other and the possibility of mis-priming with other templates.
Advantages of multiplex PCR
False negative reactions are often revealed because each amplicon provides an internal control for the other amplified fragments.
Figure 12.15 Multiplex PCR
The expense of reagents and preparation time is less in multiplex PCR.
The exponential amplification and the internal standards of multiplex PCR can be used to assess the amount of a particular template in a sample.
Applications of multiplex PCR
- Pathogen identification,
- High throughput SNP genotyping,
- Mutation analysis,
- Gene deletion analysis,
- Template quantitation,
- Linkage analysis,
- RNA detection and
- Forensic studies.
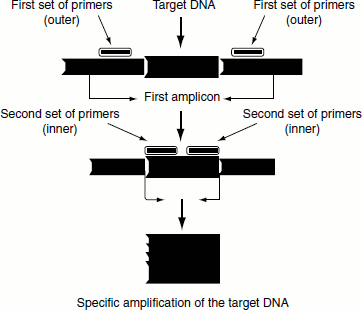
Nested PCR
This type of PCR differs from other types in that two pairs of PCR primers are used to amplify a fragment (Figure 12.16). The first pair of PCR primers amplify a fragment that is similar to a standard PCR. However, a second pair of primers called nested primers (as they are nested within the first fragment) bind inside the first PCR product fragment to allow the amplification of a second PCR product, which is shorter than the first one. The advantage of nested PCR is that if the wrong PCR fragment was amplified, the probability is quite low that the region would be amplified a second time by the second set of primers. Thus, nested PCR is a very specific PCR amplification.
Figure 12.16 Nested PCR