The genetic dictionary of the mRNA codons reveals the following important features of triplet codons:
- Degeneracy: There are 64 different triplet codons but only 20 amino acids. This proves that some amino acids must be specified by more than one codon. For example, the three amino acids arginine, serine and leucine, each have six synonymous codons. The first two bases of the synonym codons are constant, whereas the third can vary; for example, all codons starting with CC specify proline (CCU, CCC, CCA and CCG) and all codons starting with AC specify threonine. This third position is known as the ‘wobble’ position of the codon. This is because though the identity of the base at the third position can wobble, the same amino acid will still be specified. This wobbling offers some protection against mutation—if a mutation occurs at the third position of a codon, there is a good chance that the same amino acid can be specified and the encoded protein does not change.
- Non-overlapping: The code is non-overlapping, meaning that no single base can take part in the formation of more than one codon. The genetic code is read in groups of three nucleotides. After reading one triplet, the ‘reading frame’ shifts over the next three letters and not just one or two. In the following example, the code would not be read GAC, ACU, CUG, UGA…
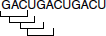 Rather, the code would be read GAC, UGA, CUG, ACU…
Rather, the code would be read GAC, UGA, CUG, ACU… 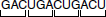
- Reading Frames: The triplet-based genetic code can be read in three possible ways. For example, the following code can be read in three different ways as:
 Each way of reading would yield completely different results. Hence, the correct way of reading the genetic code becomes a must. The genetic code is read continuously from the 5′ → 3′ direction.
Each way of reading would yield completely different results. Hence, the correct way of reading the genetic code becomes a must. The genetic code is read continuously from the 5′ → 3′ direction. - Ambiguity: The genetic code is non-ambiguous, that is, each codon specifies a particular amino acid, and only one amino acid. In other words, the codon ACG codes for the amino acid threonine, and only threonine.
- Commaless: The genetic code is commaless, which means that no codon is reserved for punctuations.
- Starting codons: AUG codon is called starting or chain initiation codon, because, it initiates the synthesis of polypeptide chain. AUG also codes for the amino acid methionine. The first AUG in the mRNA signals for translation to begin. The subsequent codons are read in the same reading frame. Translation continues until a stop codon is encountered.
- Stop codons: There are three stop codons: ‘UAA’, ‘UAG’ and ‘UGA’. The UAA is also called ochre and UAG is also called amber. These codons do not specify any amino acid and hence they are also called ‘non-sense codons’. They are also called termination codons. A reading frame between a start codon and an in-frame stop codon is called an ‘open reading frame’ (ORF).For example, consider the following sequence: 5′-GUCCCGUGAUGCCGAGUUGGAGUCGAUAACUCAGAAU-3′ The code is read in the 5′ → 3′ direction. The first AUG read in that direction sets the reading frame, subsequent codons are read in frame, until the stop codon, UAA, is reached.
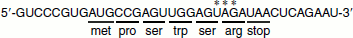 It is an absolute requirement that the codes are read in frame. In the above example, the three nucleotides marked with asterisk would specify the stop codon UAG if not read in frame.In the above sequence, there are nucleotides at either end that are outside of the ORF. Because they are outside of the ORF, these nucleotides are not used to code for amino acids and are called untranslated regions. The region at the 5′-end that is not translated is called the 5′ untranslated region or 5′-UTR. The region at the 3′-end is called the 3′-UTR. These sequences contain regulatory sequences that can regulate the gene expression.
It is an absolute requirement that the codes are read in frame. In the above example, the three nucleotides marked with asterisk would specify the stop codon UAG if not read in frame.In the above sequence, there are nucleotides at either end that are outside of the ORF. Because they are outside of the ORF, these nucleotides are not used to code for amino acids and are called untranslated regions. The region at the 5′-end that is not translated is called the 5′ untranslated region or 5′-UTR. The region at the 3′-end is called the 3′-UTR. These sequences contain regulatory sequences that can regulate the gene expression. - Universality: The genetic code has been found to be universal, because the same code applies in all kinds of living systems.
The Mitochondrial Genetic Code
Human mitochondrial DNA encodes only 22 tRNA that are used for the translation of mitochondrial mRNAs. The U of the anticodon in tRNA can pair with any of the four bases in the third codon position of the mRNA. This enables four codons to be recognized by a single tRNA. Moreover, some codons specify different amino acids in mitochondria than in the universal code.
Differences between the universal and mitochondrial genetic codes
The genomes of prokaryotic and eukaryotic cells have less genetic code variations. Among the lower eukaryotes, certain ciliated protozoans (Tetrahymena and Paramecium) use UAA and UGA as glutamine codons rather than stop codons (Table 5.1). Mycoplasma, for example, uses the stop codon UGA to specify Trp. Some UGA codons in both prokaryotes and eukaryotes (including humans) are used to specify selenocysteine. In several species of Archaea and bacteria, pyrrolysine amino acid is encoded by UAG. How the translation machinery knows when it encounters UAG whether to insert a tRNA with pyr-rolysine or to stop translation is not yet known.
Table 5.1 Universal and mitochondrial genetic codes
| Codon | Universal code | Human mitochondrial code |
|---|---|---|
| UGA | Stop | Trp |
| AGA | Arg | Stop |
| AGG | Arg | Stop |
| AUA | Ile | Met |