Nirenberg and Khorana Experiment
Marshall W. Nirenberg and Heinrich J. Matthaei (1962) made their own simple and artificial mRNA and identified the polypeptide product that was encoded by it. They used the enzyme polynucleotide phosphorylase, which randomly polymerizes any RNA nucleotides that it finds. They began with the simplest codes possible. Polynucleotide phosphorylase was added to a solution of pure uracil(U). Poly(U) polymer was created. These molecules were known as poly(U) RNAs. These poly(U) RNAs were added to 20 tubes containing all the components required for protein synthesis such as ribosomes, activating enzymes, tRNAs and other factors. Each tube contained one of the 20 amino acids, which were radioactively labelled. Of the 20 tubes, 19 tubes did not form the protein. Only one tube, the one that had been loaded with the labelled amino acid phenylalanine, yielded a product. Assuming triplet code, Nirenberg and Matthaei, therefore, found that the UUU codon could codes for the amino acid phenylalanine. Similar experiments conducted using poly(C) and poly(A) RNAs revealed that CCC encodes the amino acid proline and lysine was encoded by the AAA codon.
In an effort to decode the other codons, Nirenberg et al. made artificial RNAs containing two or three different bases. As previously mentioned, polynucleotide phosphorylase joins nucleotides randomly; as a result, these artificial RNAs contained random mixtures of the bases in proportion to the amounts of bases mixed. For example, when A and C were mixed with polynucleotide phosphorylase, the resulting RNA molecules contained eight different triplet codons: AAA, AAC, ACC, ACA, CAA, CCA, CAC and CCC. These eight random poly(AC) RNAs produced proteins containing only six amino acids: asparagine, glutamine, histidine, lysine, proline and threonine. Previous experiments had already revealed that CCC and AAA code for proline and lysine, respectively. Thus, it was concluded that the four newly incorporated amino acids could only be encoded by AAC, ACC, ACA, CAA, CCA and/or CAC.
In 1965, H. Gobind Khorana and his colleagues used another method to further decipher the genetic code. These researchers used chemically synthesized RNA molecules of known repeating sequences rather than random sequences. For example, an artificial mRNA of alternating guanine and uracil nucleotides (GUGUGUGUGUGU). This mRNA upon translation is read as two alternating codons, GUG and UGU, and encodes a protein of two alternating amino acids, cysteine and valine respectively. However, this technique could not determine whether GUG or UGU encoded cysteine.
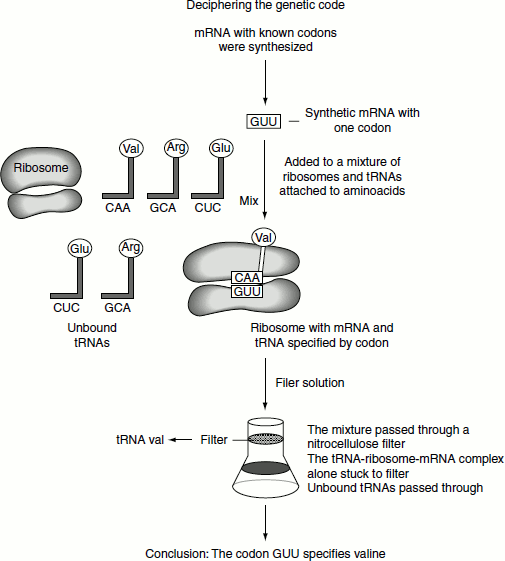
Nirenberg and Philip Leder developed a technique using ribosome-bound transfer RNAs (tRNAs). They showed that a short mRNA sequence—even a single codon (three bases)—could still bind to a ribosome, even if this short sequence was incapable of directing protein synthesis. The ribosome-bound codon could then base pair with a particular tRNA that carried the amino acid specified by the codon. They synthesized many short mRNAs with known codons. The mRNAs were then added one by one to a mix of ribosomes and aminoacyl-tRNAs with one amino acid radioactively labelled. For each reaction, they determined whether the aminoacyl-tRNA was bound to the short mRNA sequence and to the ribosome. By this method, they identified the particular aminoacyl-tRNA that was bound to each mRNA codon.