The chromosomes in eukaryotes have much complex structure than prokaryotic chromosomes. The duplication of the chromosomes of eukaryotes involves not only the replication of their DNA but also the synthesis of the associated histone and non-histone chromosomal proteins. However, at the molecular level, the replication of DNA in eukaryotes is quite similar to prokaryotes regardless of the complexity of its genome. Eukaryotic replication is semi-discontinuous and semi-conservative. Like DNA replication in E. coli, eukaryotic DNA replication occurs bidirectionally from RNA primers made by a primase; the synthesis of the leading strand is continuous, while the synthesis of the lagging strand is discontinuous. In contrast to the situation in E. coli, however, two distinct DNA polymerases, α and either δ or ε, function at the eukaryotic growing fork.
Cell Cycle
The cell cycle represents the general sequence of events that occur during the lifetime of a eukaryotic cell. The cell cycle comprises the following phases (Figure 3.9).
Figure 3.9 Cell cycle
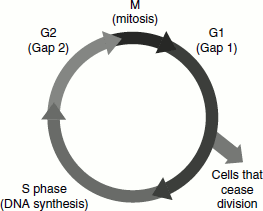
Mitosis and cell division occur during the relatively brief M-phase.
Followed by G1-phase, the longest part of the cell cycle. G1-phase gives way to the S-phase the only period in the cell cycle when DNA is synthesized.
During G2-phase, the new tetraploid cell prepares for mitosis.
Cell cycle for cells in culture occupies 16-24 h period. Cell cycle time varies for different types of cells of multicellular organism and may vary from as little as 8 h to ξ100 days. Most of this variation occurs in the G1-phase. Many terminally differentiated cells such as neurons or muscle cells never divide. They assume quiescent state known as G0-phase. Cells irreversible decision to proliferate is made during G1-phase. Quiescence is maintained, for example, if nutrients are in short supply or the cell is in contact with other cells (contact inhibition). DNA synthesis may be induced by various agents such as carcinogens or tumour viruses or by proteins known as mitogens, which bind to the cell receptor and induce cell division. The progression of the cell through the cell cycle is regulated by proteins known as cyclins and cyclin-dependent protein kinase.
Cell Cycle Control of DNA Replication
In cycling somatic cells, cells synthesize RNA and proteins during the G1-phase, preparing for DNA synthesis and chromosome replication during the S-phase. After progression through the G2-phase, cells begin the complicated process of mitosis. The concentrations of cyclins the regulatory subunits of heterodimeric protein kinases that control cell cycle events, increase and decrease as cells progress through the cell cycle. Cyclin kinases have no kinase activity unless they are associated with cyclin. Each cyclin-dependant kinase (CDK) can associate with different cyclins and the type of associated cyclin determines which proteins are phosphorylated by a particular cyclin–CDK complex.
There are three major classes of cyclin–CDK complex; namely, G1 cyclin–CDK complex, S cyclin–CDK complex and mitotic cyclin–CDK complex. When cells are stimulated to replicate, G1 cyclin–CDK complex are expressed first. These prepare the cell for the S-phase by activating transcription factors that promote the transcription of genes encoding enzymes required for DNA synthesis and the genes encoding S-phase cyclin–CDKs. The three G1 cyclins associate with CDK to form S-phase-promoting factor (SPF). SPF phosphorylates and regulates proteins required for DNA synthesis. The degradation of S-phase inhibitor triggers DNA replication.
The S-phase cyclin–CDK complexes, Clb 5-CDk and Clb 6-CDK begin to accumulate in G1; however, they are inactivated by the binding of an inhibitor, called Sic1, that is expressed late in mitosis and early G1. Sic1 specifically inhibit B-type cyclin–CDK complexes, but has no effect on G1 cyclin CDK complexes, i.e., it functions as an S-phase inhibitor.
Sic 1 inhibitor is degraded following its polyubiquitination by a ubiquitin ligase called SCF. Once Sic1 is degraded, the S-phase cyclin–CDK complex induces replication by phosphorylating multiple proteins bound to replication origins (Figure 3.10).
Figure 3.10 Cyclins regulating initiation of DNA replication
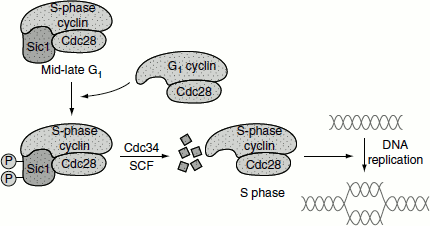
Eukaryotic Replication Origins
Eukaryotic chromosomes are replicated from multiple origins (Figure 3.11). Some of these initiate replication early in the S-phase, some later and still others towards the end. However, no eukaryotic origin initiate more than once per S-phase. Moreover, S-phase continues until replication from all origins along the length of each chromosome results in the replication of the chromosomal DNA in its entirety. These two factors ensure that the correct gene copy number is maintained as cell proliferates.
Figure 3.11 Eukaryotic origins of replication
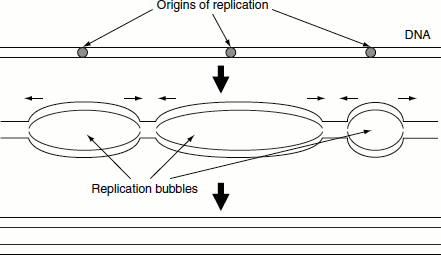
Initiation of DNA Replication
Yeast replication origins contain 11 bp conserved sequence to which is bound a hexameric protein, the origin recognition complex (ORC). ORC remains associated with origin during all phases of the cell cycle. Several replication factors are required for the initiation of DNA replication such as Cdc 6, Cdt 1, MCM 10 and MCM hexamer a complex of six additional closely related MCM proteins. These proteins associate with origins during G1 or M. During G1, the various initiation factors assemble with ORC into a pre-replication complex at each origin.
The initiation of replication requires an active S-phase cyclin–CDK complex, a heterodimeric protein kinase Dbf 4-Cdc 7, which is expressed in G1. Dbf 4-dependent kinase, Cdc 7, is often called DDK. DDK phosphorylates the components of pre-replication complex. This leads to the binding of Cdc 45 which is followed by the activation of MCM helicase that unwinds the parental DNA strand and the release of the phosphorylated Cdc 6 and Cdt 1 initiation factors. RPA binds to the resulting single-stranded DNA. A complex of DNA polymerase a and primase initiates the synthesis of the daughter strands. ORC remains bound to the origin sequence in the daughter double-stranded DNA but the phosphorylated initiation factors cannot assemble the pre-replication complex on it. B-type cyclin CDK complexes maintain the initiation factors in a phosphorylated state through the remainder of S-phase, G2-phase and early anaphase. The initiation factors cannot assemble pre-replication complex until B-type cyclins are degraded following their polyubiquitination in the late anaphase (Figure 3.12).
Figure 3.12 Eukaryotic replication-initiation and elongation
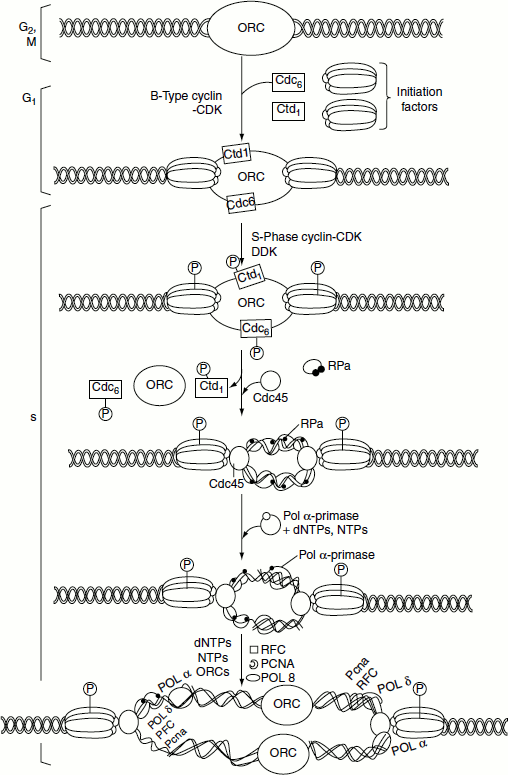
Elongation
The DNA polymerase α and primase initiate the synthesis of the daughter strands, however, DNA polymerase α is not capable of a lengthy DNA synthesis, as it has low processivity. This is followed by PCNA binding that aids the binding of DNA polymerase δ, which carries out the rest of the synthesis.
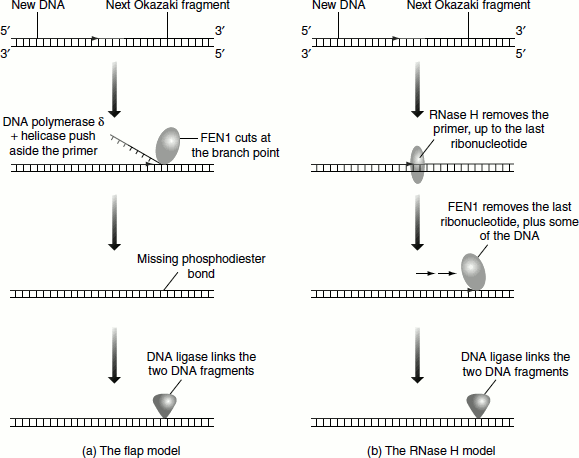
Flap endonuclease, FEN1 (previously called MF1), is associated with DNA polymerase 8 complex at the 3′-end of an Okazaki fragment in order to degrade the primer from the 5′-end of the adjacent fragment. FEN1 cannot initiate the primer degradation because the ribonucleotide at the extreme 5′-end of the RNA primer carries a triphosphate that blocks FEN1 activity. The alternative models to circumvent this problem have been proposed (Figure 3.13).
The first possibility is that a helicase breaks the base pairs holding the primer to the template strand, enabling the primer to be pushed aside by DNA polymerase δ, as it extends the adjacent Okazaki fragment into the region thus exposed. The flap that results can be cut off by FEN1. Alternatively, most of the RNA component of the primer could be removed by RNase H, which can degrade the RNA part, i.e., the primer. However, this enzyme cannot cleave the phosphodiester bond between the last ribonucleotide and the first deoxyribonucleotide. This ribonucleotide will carry a 5′-monophosphate than a triphosphate and hence can be removed by FEN1.
Figure 3.13 DNA elongation assisted by FEN1
Licensing of DNA Replication
Various chromosomal regions are not replicated simultaneously. Clusters of 20–80 adjacent replicons are activated. Replicons are activated throughout the S-phase until the entire chromosome has been replicated. During this process, the replicons that have already been replicated are distinguished from those that are not, i.e., cells chromosomal DNA is replicated only once per cell cycle. A pre-replication complex is assembled at the origin only during the G1-phase of the cell cycle. G1 is the only period during which pre-replication complex can form. Hence, this process is known as licensing. However, licensed pre-replication complex cannot initiate the replication until it is activated during the S-phase.
Termination of Replication
Eukaryotic DNAs are linear. The ends of linear chromosome cannot be replicated easily. This is because the RNA primers at the 5′-end of a completed lagging strand cannot be replaced with DNA, as the polymerase required to do this would have no place to bind. At the extreme end of a chromosome, there is no way to synthesize this region when the last primer is removed. Therefore, the lagging strand is always shorter than its template by at least the length of the primer. This is the so-called ‘end-replication problem ′. For this reason, eukaryotic chromosomes have linear DNA sequences at the ends called telomeres.
Telomeres and Telomerases
Telomeric DNA has an unusual sequence. It consists of up to several thousand tandem repeats of a simple, species-dependent and G-rich sequence concluding the 3′ terminus of each chromosomal terminus. The enzyme that synthesizes the G-rich strand of telomeric DNA is named telomerase. Telomerase adds tandem repeats of the telomeric sequence TTTGGG to the 3′-end of the lagging-strand template.
Figure 3.14 Replicating ends of eukaryotic DNA by telomerases
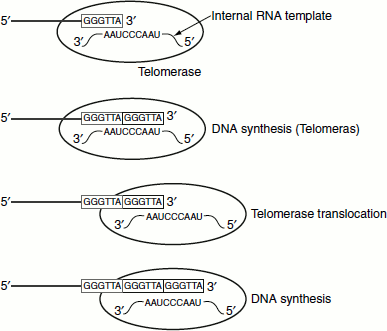
Telomerase is a ribonucleoprotein. Its RNA component contains a segment that is complementary to the telomeric sequence. This sequence acts as a template in a kind of reverse transcriptase reaction that synthesizes the telomeric sequence, translocates to the DNA’s new 3′-end and repeats the process (Figure 3.14).
Without the action of telomerase, a chromosome would be shortened by 50–1,000 nucleotides with every cycle of DNA replication and cell division. The loss of telomerase function in somatic cells is the basis for aging in multicellular organisms. Cancer cells have active telomerases.
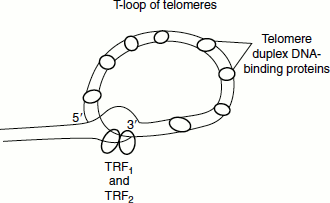
Exposed telomeric DNA would result in the end-to-end fusion of chromosomes, which is a process that will lead to chromosome instability and eventually cell death. This is prevented by capping the DNA. Telomeric DNA is specifically bound by proteins that sequester the DNA ends forming T-loops.