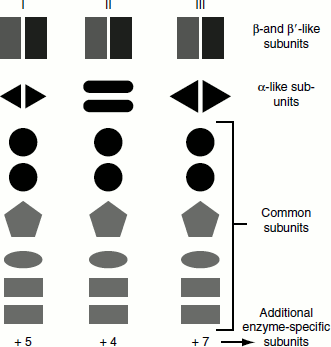
Eukaryotic cells contain three different RNA polymerases, designated as I, II and III, which catalyse the transcription of genes encoding different classes of RNA. The subunit structure of yeast RNA polymerase is depicted in the following (Figure 4.11).
Figure 4.11 Yeast RNA polymerases and their submits
RNA Polymerase-I
RNA polymerase-I (RNA Pol-I) is located in the nucleolus and synthesizes the precursor RNA (Pre-rRNA), which is processed into the 28S, 5.8S and 18S rRNAs. It accounts for nearly half the total number of RNA found in the cell and is insensitive to α-amanitin, a toxin from the mushroom called Amanita phalloides. The action of the toxin on the three classes of RNA polymerase activity varies and this characteristic is used to distinguish the polymerases. The enzyme is tightly regulated, so that the ribosomal RNA synthesis keeps pace with the cells’ protein requirements for growth, development and division. The holoenzyme includes two large polypeptide subunits and 4 to 10 smaller subunits, which varies according to the source. Some of these smaller subunits are common to the other two polymerases. RNA Pol-I requires at least two transcriptional factors for activity. The transcription factors help in the binding of promoter and in the initiation of transcription.
RNA Polymerase-II
The enzyme RNA polymerase-II (RNA Pol-II) transcribes the pre-mRNA gene of the cell and is thus responsible for the transcription of the largest part of the genome. RNA Pol-II also produces four small RNAs that take part in RNA Splicing [U1, U2, U3 and U4]. It present in nucleoplasm. It is very sensitive to the mushroom poison α-amanitin.
Of the three polymerases, RNA Pol-II is the most intensively studied. This polymerase recognizes three different elements of a gene.
- A selector sequence containing a TATA box and a short sequence.
- An upstream promoter sequence
- An enhancer sequence that may be located at different site in different genes
Seven transcription factors assist the RNA Pol-II for a specific binding of the enzyme to the DNA promoter and to initiate transcription. The enzyme has 12 subunits namely RPB1 to RPB12. The three largest subunits, i.e., RPB1, RPB2 and RPB3, constitute the core of the enzyme. RPB1 and RPB2 are functionally comparable toβ and β′ of the E. coli RNA polymerase. RPB3 has been proposed to be analogous to the α-subunit of the E. coli polymerase. The subunits RPB4 and RPB7 may contribute to fine-tuning the transcription apparatus. The subunits RPB5, RPB6, RPB8, RPB10 and RPB12 are common subunits and are found even in RNA Pol-I and RNA Pol-II.
The largest subunit of RNA Pol-II has a CTD (carboxyl terminal domain) consisting of multiple repeats of a heptamer. The CTD is involved in the regulation of the initiation of the transcription. Phosphorylation of the serine/threonine residues of the heptanucleotide repeat of CTD is required for promoter clearance and for proceeding to the transcription elongation.
RNA Polymerase-III
The enzyme RNA polymerase-III (RNA Pol-III) is present in nucleoplasm. It transcribes the following genes.
- The tRNA genes;
- The 5S rRNA genes that is found in the large ribosomal subunit (60S) of eukaryotes;
- The genes of the RNA (e.g., Usn RNAs) that assist in the processing of pre-RNAs by spliceo-some and
- In addition, the genes of 7S RNA of the signal recognition particle (SRP), which is involved in the transport of proteins into the endoplasmic reticulum.
It is the most structurally complex RNA polymerase. In yeast, the complete molecule is made up of 14 distinct subunits. Like other enzymes, it has two large subunits associated with smaller subunits. A few smaller subunits are common to the other polymerase. It is less sensitive to α-amanitin.
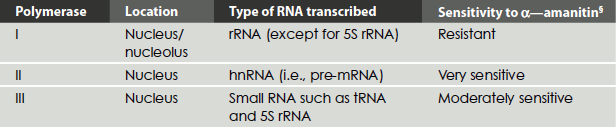
Table 4.4 describes the properties of three types of eukaryotic RNA polymerases.
Template-independent RNA Polymerases
Apart from the three major classes of RNA polymerases, there are a few RNA polymerases that are found in cells that do not require template for polymerization, but they require pre-existing RNA. These include:
- tRNA-specific nucleotidyl transferase, which adds CCA sequence to 3′-end of tRNA during the post-transcriptional modification of pre-tRNA.
- Poly(A) polymerase, which adds poly(A) tail to 3′-end of hnRNA during the post-transcriptional modification of eukaryotic pre-mRNA.
Table 4.4 Eukaryotic RNA polymerases

Mechanism of Action of RNA Pol-II
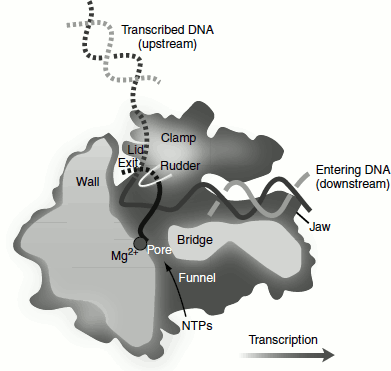
Roger Kornberg determined the X-ray structure of the yeast RNA Pol-II. RNA Pol-II has two Mg2+ ions at its active site. Two proteins of RNA Pol-II, namely Rpbl and Rpb2, named the clamp forms a cleft in which DNA is trapped. This accounts for the enzymes infinite processivity. DNA unwinds by three bases before entering the active site, which is contained on Rpbl. A portion of the Rpb2 called the ‘wall’ directs the template out of the ‘cleft’ in an approximately 90° turn. Consequently, the template base at the active site (+1) points towards the floor of the cleft where it can be read out by the active site. This base is paired with the ribonucleotide at the 3′-end of the RNA, which is positioned above a ‘pore’ at the end of a ‘funnel’ through which NTP’s presumably gain access to the otherwise sealed off-active site. The RNA-DNA hybrid adopts a non-standard conformation intermediate between those of A and B DNAs. After about one turn of hybrid helix forms, a loop extending from the clamp called the ‘rudder’ separates the RNA and template DNA strands, thereby permitting the DNA double helix to reform as it exits the enzyme. The highly conserved helical segment of rpb1, called the ‘Bridge’ (since it bridges the two pincers forming the enzyme cleft), non-specifically contacts the DNA template base at +1 position. The bridge alternates between two conformations straight and bent. This makes a movement of about 3 to 4 Å. Translocation of RNA–DNA assembly occurs through the bending of the bridge helix so as to push the paired nucleotides at position +1 to position −1. The recovery of the bridge helix to its straight conformation would yield an empty site at position +1 for the entry of the next NTP, thereby preparing the enzyme for a new round of nucleotide addition (Figure 4.12).
Figure 4.12 RNA polymerase II catalyzing transcription