Protein synthesis inhibitors serve two major purposes (Figure 6.25) (Table 6.1).
First, they have been very useful scientifically in elucidating the biochemical mechanisms of protein synthesis.
Second, some of these inhibitors affect prokaryotic but not eukaryotic protein synthesis and thus are medically important antibiotics.
Streptomycin
Streptomycin is an aminoglycoside antibiotic that affects the function of the prokaryotic 30S sub-unit. Low concentrations of streptomycin induce mRNA misreading. As the result, improper amino acids are incorporated into the polypeptide. Codons with pyrimidines in the first and second positions are particularly susceptible to streptomycin-induced misreading. These reading errors are not frame shift mistakes; therefore, totally aberrant proteins are not made at low streptomycin levels. Thus, susceptible cells are not killed, but their growth rate is severely decreased. At high concentrations of streptomycin, non-productive 70S ribosome–mRNA complexes accumulate, preventing the formation of active initiation complexes with new mRNA.
Table 6.1 Some of the protein synthesis inhibitors and their action
| Inhibitor | Mode of action | Organism inhibited |
|---|---|---|
| INHIBITORS OF INITIATION | ||
| Aurintricarboxylic acid | Inhibits initiation factor binding to the 30S subunit | Prokaryotes |
| Kasugamycin | Inhibits f-Met-tRNAf Met binding | Prokaryotes |
| Streptomycin | Inhibits formation of initiation complexes | Prokaryotes |
| INHIBITORS OF ELONGATION | ||
| Tetracycline | Inhibits aminoacyl-tRNA binding at A site | Prokaryotes |
| Streptomycin | Leads to codon misreading, insertion of improper amino acid | Prokaryotes |
| Sparsomycin | Peptidyl transferase inhibitor | Prokaryotes |
| Chloramphenicol | Blocks peptidyl transferase activity by binding to 50S subunit | Prokaryotes |
| Erythromycin | Blocks peptidyl transferase activity by binding to 50S subunit | Prokaryotes |
| Cycloheximide | Inhibits translocation of peptidyl-tRNA | Eukaryotes |
| Fusidic acid | Inhibits EF-G:GDP dissociation from ribosome | Both prokaryotes and eukaryotes |
| Thiostrepton | Inhibits ribosome-dependent EF-Tu and EF-G GTPase | Prokaryotes |
| Diphtheria toxin | ADP-ribosylates and inactivates eEF2 | Eukaryotes |
| INHIBITORS THAT BRINGS ABOUT PREMATURE TERMINATION | ||
| Puromycin | Aminoacyl-tRNA analogue, acts as a peptidyl acceptor and aborts further peptide elongation | Both prokaryotes and eukaryotes |
| Ricin | Inactivates 28S rRNA | Eukaryotes |
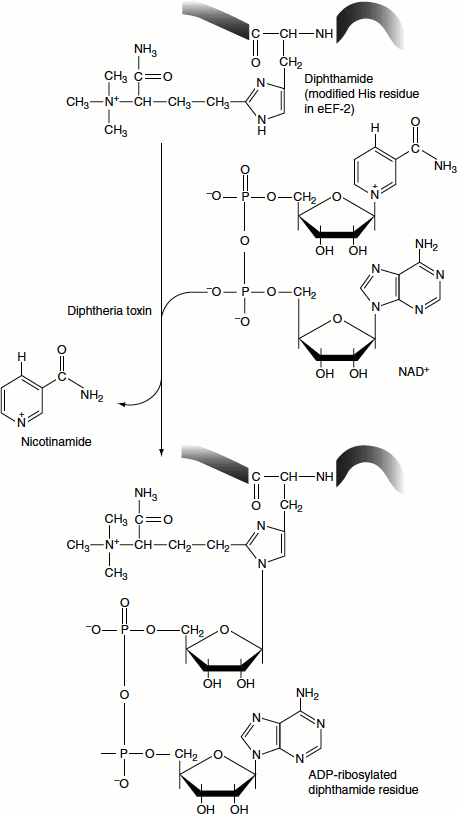
Figure 6.25 Inhibitors of protein synthesis
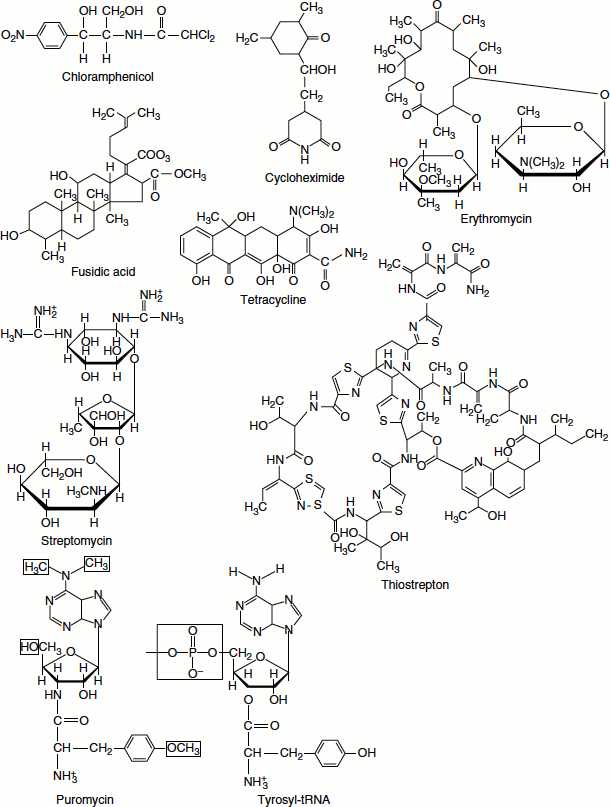
Figure 6.26 ADP-Ribosylation of the diphthamide moiety of eukaryotic EF-2
Puromycin
Puromycin structurally resembles 3′-end of aminoacyl-tRNAs. Puromycin binds at the A site of both prokaryotic and eukaryotic ribosomes. The binding is not dependent on EF-Tu (or EF1). Puromycin acts as an acceptor of the peptidyl chain from peptidyl-tRNA
Diphtheria Toxin
Diphtheria arises from infection by Corynebacterium diphtheriae, a bacterium carrying bacteriophage corynephage. Diphtheria toxin is a phage-encoded enzyme secreted by these bacteria. The toxin is capable of inactivating a number of GTP-dependent enzymes. Diphtheria toxin is an NAD+-dependent ADP-ribosylase. It covalently attaches to an ADP-ribosyl moiety derived from NAD+.
One target of diphtheria toxin is the eukaryotic translocation factor, EF2, which has a modified His residue known as diphthamide. Diphthamide is generated post-translationally on EF2; its biological function is unknown. (EF-G of prokaryotes lacks this unusual modification and is not susceptible to diphtheria toxin.) Diphtheria toxin specifically ADP-ribosylates an imidazole-N within the diphthamide moiety of EF2 (Figure 6.26).
ADP-ribosylated EF2 can still bind GTP but cannot function in protein synthesis. As diphtheria toxin is an enzyme and can act catalytically to modify many molecules of its target protein, just a few micrograms sufficient to cause death.
Diphtheria toxin catalyses the NAD+-dependent ADP-ribosylation of selected proteins like ADP-ribosylation of the diphthamide moiety of eukaryotic EF2. (Diphthamide=2-[3-carboxamido-3-(trimethylammonio)propyl]histidine.)
Ricin
Ricin is an extremely toxic glycoprotein produced by the plant Ricinus communis (castor bean). The protein is disulphide-linked. It is a heterodimer of roughly equal 30-kDA subunits namely A and B.
The A-subunit (32 kDA) is an enzyme and is the toxic subunit; ricin gains entry to cells with the help of the B-subunit (33 kDA) which is a lectin. (Lectins form a class of proteins that bind to specific carbohydrate moieties commonly displayed by glycoproteins and glycolipids on cell surfaces.)
Endocytosis of bound ricin followed by disulphide reduction releases the A chain. The A chain enters the cytosol and catalytically inactivates eukaryotic large ribosomal subunits. A single molecule of ricin A chain in the cytosol can inactivate 50,000 ribosomes and can kill a eukaryotic cell.
Ricin A chain specifically attacks a single and highly conserved adenosine (an A at position 4,256) in the eukaryotic 28S rRNA. Ricin A chain has an N-glycosidase activity that removes the adenine base, leaving the rRNA sugar–phosphate backbone intact. The removal of this single base is sufficient to inactivate a 60S large subunit. The adenine in this highly conserved region of the 28S rRNA sequence is believed to be crucial to the functions of the 60S subunit that involve EF1 and EF2.