S1 Nuclease
It is an endonuclease, which cleaves single-stranded DNA or single-strand protrusion of double-stranded DNA with cohesive ends. Because of S1 nuclease action, cohesive ends are converted into blunt ends. Thus, S1 nuclease is used to remove the incompatible ends.
DNA Pol-I, Klenow fragment
This fragment has the polymerase activity and 3′ → 5′ exonuclease activity of DNA Pol-I but does not have the 5′ → 3′ exonuclease activity. This 5′ → 3′ exonuclease activity is often troublesome because it degrades the 5′ terminus of primers that are bound to DNA templates and removes 5′-PO4 from the terminal DNA fragments that are to be used as substrates for ligation. The Klenow fragment can be used in the end filling and synthesis of DNA in cDNA clone.
Alkaline phosphatase
The cohesive ends of restriction enzyme treated plasmids instead of joining with the target DNA, sometimes reseal without taking the insert (target DNA) and are recircularized. To overcome this problem, the restricted vector (plasmid) is treated with the enzyme alkaline phosphatase, which removes the terminal 5′-PO4 group. The restriction fragments of the target DNA to be cloned are not treated with alkaline phosphatase. Therefore, the 5′end of the target DNA can covalently join with the 3′-end of the plasmid. Ligase action completes the formation of the rDNA.
Reverse transcriptase
Reverse transcriptase is used in the synthesis of cDNA using RNA template and also for the construction of cDNA clone bank.
Deoxyribonuclease I (DNAse I)
It is an endonuclease, which digests single- and double-stranded DNAs. This enzyme is useful for a variety of applications including nick translation, DNA foot printing, bisulphite-mediated mutagenesis and RNA purification. These enzymes have a role in genetic engineering and thus produce required specifications.
CLONING VECTORS
A DNA molecule that carries foreign DNA into a host cell, replicates inside the host cell and produces many copies of itself is called a cloning vector. Cloning is frequently employed to amplify DNA fragments containing genes.
Some essential characteristics of a cloning vector are:
- It contains a sequence that allows for the propagation of itself in the host.
- It contains an insertion site for the foreign DNA—also called an MCS (multiple cloning site) that can be cut by several restriction enzymes.
- It contains specific control systems such as promoters, terminators and ribosome-binding site, so that the cloned DNA is expressed properly.
- It contains marker genes that allow for the selection of the host cells that contain the insert (DNA of interest).
Types of Cloning Vectors
The various types of cloning vectors used in rDNA technology include:
- Plasmids vectors,
- Bacteriophage vectors,
- Cosmid vectors,
- Shuttle vectors,
- Yeast episomal plasmid (YEP vectors),
- Yeast artificial chromosomes (YAC vectors) and
- Ti plasmids.
Plasmid vectors
Plasmids are found in bacteria and are autonomously replicating extrachromosomal circular DNA molecules. They are small between 2 and 8 Kb and often have a high copy number. They also encode antibiotic-resistant gene for propagation in a suitable host and contain unique restriction enzyme sites for cloning purposes. Plasmids are frequently used, as they are the easiest class of vector to work with, as their isolation from bacteria, modification with enzymes, ligation and introduction into the host are quite easier.
One of the earliest plasmid vectors was pBR322 that was named after Bolivar and Rodriguez who were involved in engineering this plasmid. pBR322 is a small plasmid of 4.36 Kb in size. It replicates extrachromosomally in the bacterium E. coli. pBR322 encodes antibiotic-resistant genes for ampicillin (Amp) and tetracycline (Ter), which facilitate selection on antibiotic-containing plates. It also contains unique restriction sites for many enzymes such as Sal I, Eco R I, Bam H I and Pst I (Figure 9.12).
Figure 9.12 Plasmid vectors
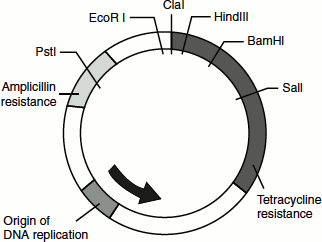
The restriction digestion of pBR322 with Sal I allows the insertion and ligation of Sal I-digested foreign DNA into the middle of the tetracycline gene. This helps in the selection of the transformed bacterial cells (cells bearing the rDNA) for Amp resistance that will be intact, followed by subsequent screening for tetracycline sensitivity that arises due to the interruption of the Ter-resistant gene because of the insert. In this way, bacterial cells bearing the rDNA can be selected and subsequently cultured, thus amplifying the gene of interest. With the advanced genetic engineering techniques, the pBR322-based plasmid vectors were now modified to be more versatile and user-friendly plasmids.
The pUC family plasmids
These plasmids were developed by Messings and co-workers at the University of California. One revolutionary feature introduced in these plasmids was the presence of MCS’s. The MCS is a short stretch of DNA that contains the recognition sequence for a large number of restriction enzymes. Thus, the choice of the restriction enzyme required to digest the vector and the target DNA is increased.
The pUC plasmids also contains lacZ′ gene as the marker gene. The lacZ′ gene codes for the α-peptide portion of the enzyme β-galactosidase. When this plasmid is inserted into an E. coli, which lacks lacZ′ segment (i.e., lacZ′ mutant), the bacterial and plasmid genes complement each other to produce a functional β-galactosidase. This process is called ‘α-complementation’. If a foreign gene is inserted into this lacZ′ gene of pUC plasmids, then it cannot complement and no functional β-galactosidase is formed (Figure 9.13).
Figure 9.13 Cloning vector-pUC19
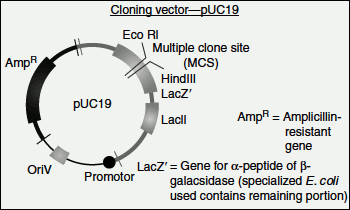
When the lacZ is expressed in a suitable host, it is capable of degrading the chromogenic substrate X-gal (5-bromo-4-chloro-3-indoyl β-D-galactopyranoside) to produce a blue colour. If the DNA of interest is introduced into the lacZ gene, the β-galactosidase protein will not be produced and hence any clone with the insert is incapable of degrading X-gal substrate. Thus, recombinant bacterial cells will appear colourless while non-recombinants will be blue in colour.
Disadvantages of plasmid vectors
- The size of the target DNA inserted into these vectors is usually limited.
- Recombinant plasmids can be extremely unstable in certain hosts; for example, some strains of Bacillus.
Bacteriophage vectors
Bacteriophages are viruses that infect bacteria. A number of different bacteriophages have been studied and well characterized. Many gene-cloning experiments are performed with bacteriophage lambda λ(X phage), which infects E. coli. The λ phages have a protein head that contains the phage DNA (approximately 49 Kb). It also has a tail and tail fibres. The tail fibres help in the adsorption of the phages onto the surface of the bacterium. Once inside the bacterial cell, the λ phage DNA enters either a lytic or lysogenic cycle.
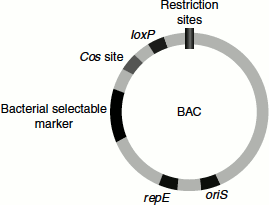
In the ‘lytic cycle’, the host cell’s machinery is utilized by the λ phages for carrying out it own molecular processes such as replication, transcription and translation, following which the phage particles are assembled. Once a threshold number of phages are assembled, the bacterial cell bursts releasing the mature phages that can infect other bacterial cells. In the ‘lysogenic cycle’, the phage that enters the bacterial cell remains associated with the host chromosome (Figure 9.14).
Figure 9.14 Lytic and lysogenic cycle of λ phage
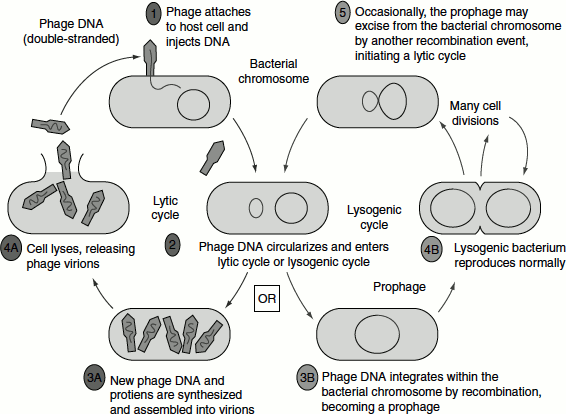
λ Phage’s DNA can be modified in several ways, thus making it suitable to be used as a cloning vector. Infection can be achieved by packaging rDNA into phage outside the natural environment by a process known as ‘in vitro packaging’.
In vitro packaging is done by isolating a combination of phage mutants that can over produce the various protein components of mature phage while lacking one of the essential proteins. These can be mixed together in the correct proportion to generate ‘packaging mixes’.
When the phage DNA is added to the packaging mix, it is packaged by a specific cleavage mechanism at a sequence termed the Cos site (cohesive site). Thus, infective phages can be generated in vitro. These are made to infect the E. coli cells. The infected cells are subcultured on agar plates. Successful infection results in the production of small lytic zones (clearings) on the agar plate, which are called ‘plaques’. The rDNA containing cells will not produce such plaques.
There are two broad classes of λ phage vectors used in gene cloning, which are ‘replacement vectors’ and ‘insertion vectors’.
Figure 9.15 Replacement vector

Lambda replacement vectors
These have been designed to accommodate large fragments of DNA. This is made possible by the removal of the DNA of the viral genome that is unnecessary for its function by restriction digestion. The removed fragment is termed the ‘stuffer’ fragment. The remaining DNA is termed the ‘arms’. The foreign DNA of interest digested with the appropriate restriction enzyme can then be used to replace the stuffer fragment by ligation to the arms. These recombinant fragments can then be used to infect bacterial cells after in vitro packaging (Figure 9.15).
Lambda insertion vectors
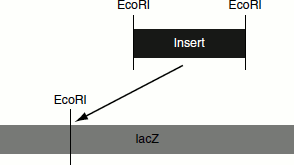
In these vectors, the DNA of interest is inserted at a single site without the removal of viral sequences. Thus, only smaller fragments can be inserted by this method when compared to the replacement vectors (Figure 9.16).
Figure 9.16 Insertion vectors
Disadvantages of bacteriophage vectors
The difficulties in manipulation; for example, isolating arms, preparation of packaging mixes, etc. are some of the disadvantages. However, the introduction of cloning kits has greatly reduced the laborious aspects of using the bacteriophage vectors.
Cosmid Vectors
Cosmids are novel vectors that combine the features of plasmid and λ phages cosmids are plasmids with cos site. They resemble plasmids in having an antibiotic-resistant marker and can replicate in bacterial cells. However, unlike plasmids, they can accommodate larger DNA fragments of up to 45 Kb. This is possible because of their cos site. The presence of the cos site allows the vector and target DNA to be packaged as if it were λ DNA. The packaged DNA is then used to infect E. coli cells. Once injected, the cosmid can replicate like a plasmid in the bacterial cell.
The various steps for cloning DNA fragments in cosmid vectors are (Figure 9.17):
- The cosmid vector is first cut with a restriction enzyme and then ligated to 35–45-Kb restriction fragments of foreign DNA with complementary cohesive ends.
- If the concentration of foreign DNA is sufficiently high, the ligation reaction generates long DNA molecules that are multiple restriction fragments of the foreign DNA separated by the cosmid DNA. These ligated molecules resemble the concatemers that form during the replication of λ phage in a host cell and can be packaged in vitro by using packaging mixes.
- In the packaging reaction, the λ Nu1 and A proteins bind to cos sites in the ligated DNA and directs the insertion of the DNA between two adjacent cos sites into empty phage heads. Packaging will occur as long as the distance between adjacent cos sites does not exceed above 50 Kb.
- Phage tails are then attached to the filled heads, producing viral particles that contain a recombinant cosmid DNA molecule.
- When these recombinant virions are plated on agar plates containing E. coli cells, they bind to phage receptors on the cell surface and inject the packaged rDNA into the bacterial cell.
- The injected DNA does not encode any λ proteins and hence no viral particles are formed in the infected cells and no plaques develop on the plate.
- The inserted DNA forms a circular plasmid or a cosmid carrying the inserted DNA fragment in each host cell.
- Like plasmids, the cosmid vector also replicates autonomously and is segregated to the daughter cells.
- Cosmid vectors containing transformed cells can be selected on antibiotic plates.
Cosmid vectors are the best to clone eukaryotic genes, as many of the eukaryotic genes are of the order of 30–40 Kb in length.
Shuttle Vectors
Certain vectors can replicate in different host systems; for example, in E. coli and in yeast. Such vectors are called shuttle vectors. The shuttle vectors carry different origins of replication that enable them to replicate in different host systems. Usually, the vector is cloned in prokaryotic systems and then the recombinant vectors are grown in eukaryotic cells. One of the most common types of shuttle vectors is the yeast shuttle vector.
Figure 9.17 Construction of a cosmid library. Cosmids are plasmid vectors that carry the cos sites from the λ phage as well as a standard plasmid origin of replication and a drug-resistance gene [here, ampicillin (ampR)]. To clone genomic DNA into a cosmid, the vector is linearized with the restriction enzyme BamHI and the genomic DNA is partially digested with Sau3AI, which leaves BamHI-compatible ends. DNA fragments ranging from 35 to 45 kilobase pairs are isolated and ligated to linearized cosmid DNA, forming tandem arrays of vector and genomic DNA fragments, A λ packaging extract recognizes and packages any ligated DNA that carries two cos sites 35 to 45 kb apart. These cosmid virions are introduced into Escherichia coli cells by infection and replicate as drug-resistant plasmids. In this way, a vector, is available. It represents the entire genome of a microrganism

Yeast Episomal Plasmids
YEPs are shuttle vectors. They can replicate in E. coli and also in yeast; for example, pJDB219 (Figure 9.18). The artificial plasmid has the following features:
Figure 9.18 Shuttle vector-YEP

- The 2-μm circular plasmid of yeast.
- The entire pBR322 sequence.
- The Leu2 gene (that acts as the selectable marker gene) of the yeast chromosome. This also carries an origin of replication.
2-μm circular plasmid
It is the plasmid found in several strains of yeast. It is 6 Kb in size and has high copy number between 70 and 200. It replicates autonomously using the enzymes provided by the host cell.
Leu2 Gene
This is a yeast chromosomal gene and encodes the enzyme isopropyl malate dehydrogenase. This is one of the enzymes involved in the conversion of pyruvic acid to leucine. When this gene is used as a selectable marker, the host yeast cell must be a mutant that has a non-functional Leu2 gene. Such cells will be unable to synthesize leucine and will grow only when this amino acid is supplemented to the growth medium (Figure 9.19).
On the other hand, cells that are transformed by the YEP containing Leu2 gene can grow in the medium lacking leucine. This enables the selection of transformed yeast cells.
The pBR322 plasmid segment of the YEP has marker genes the ampr and the terr genes, a bacterial origin of replication that will enable the YEP to replicate in a bacterial system too.
Figure 9.19 Selection of transformed cells using Leu2 gene as a selectable marker
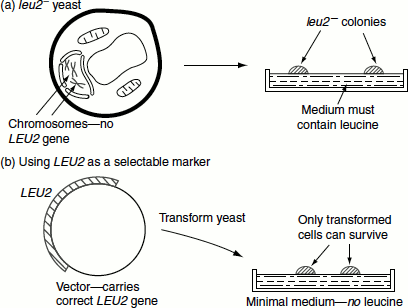
Figure 9.20 Recombinant YEP13 molecules

The YEP vector is episomal in nature and can co-integrate with one of the yeast chromosomes. Integration is made possible by homologous recombination between the Leu2 gene and the yeast mutant Leu2 gene. The plasmid portion of the vector may remain integrated or it may be excised later (Figure 9.20).
The initial cloning experiments are done with E. coli cells. The recombinants are selected. The recombinant plasmids are then purified and introduced into yeast where the new gene will express.
YAC Vectors
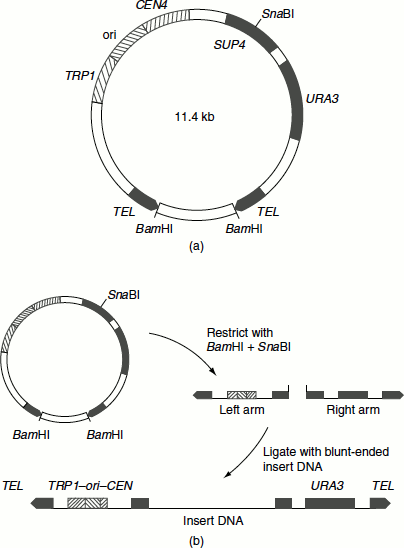
Large DNA segments can be cloned in YAC. They are linear segments that contain all molecular components required for replication in yeast namely (Figure 9.21(a)):
- A replication origin known as autonomously replicating sequence (ARE).
- A centromere.
- The telomeres.
They are called mini chromosomes. The DNAs of several hundred Kb (200–400 Kb) can be introduced into YACs and successfully cloned (Figure 9.21(b)).
Expression vectors
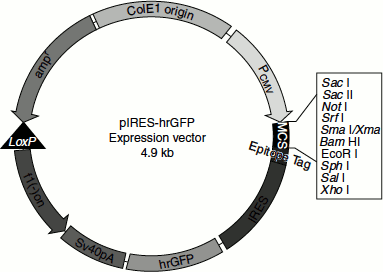
For a eukaryotic gene to be expressed in E. coli (in bacterial system), the gene must have a promoter, terminator, ribosome-binding site and other regulatory signals recognized by the bacteria. Bacterial RNA polymerase cannot recognize eukaryotic promoter, etc. Therefore, vectors are designed in such way that the inserted foreign gene is placed under the control of E. coli expression signals. Such vectors are called expression vectors since they allow the expression of a foreign gene.
Figure 9.21(a) YAC (b) The cloning strategy with pYAC3
An expression vector should carry a strong promoter, so that the cloned gene is transcribed at the highest rate. If the recombinant protein produced harmful effects on the host, then its synthesis must be regulated to prevent toxic levels. Some of the promoters used in expression vectors include E. coli Lac promoter, E. coli trp promoter, E. coli tac promoter (a hybrid of lac and trp promoters; stronger than lac or trp and is induced by IPTG), λPL promoter, etc. These promoters can be recognized by E. coli RNA polymerase.
The efficiency of translation of an mRNA in bacteria is dependent on the presence of a ribosome-binding site (i.e., Shine-Dalgarno sequence) and also on the distance between this site and the first AUG codon. Therefore, to express a eukaryotic gene, the Shine-Dalgarno sequence is usually included in the expression vector. The initiation codon of the eukaryotic gene should be placed downstream at the correct distance from the Shine-Dalgarno sequence (Figure 9.22). The expression vectors so designed when expressed will give a ‘hybrid protein’ containing a few amino acids from the prokaryotic protein and the remaining from the eukaryotic protein (i.e., inserted gene). This is called a ‘fusion protein’. Such fusion proteins are more stable in bacteria and are not degraded by bacterial proteases.
Figure 9.22 Expression vector
Vectors for Plants
Ti plasmid
The soil-borne bacteria Agrobacterium tumefaciens and A. rhizogenes causes crown gall disease and hairy root disease on the stems of numerous plants. A tumour-inducing plasmid from these bacteria is used by scientists to insert desirable genes into plant chromosomes. This plasmid is called Ti plasmid (as it induces tumour).

The size of the Ti plasmid ranges between 180 and 250 Kb. It contains a T-DNA region of about 23–25 Kb. Ti plasmid also contains regions for opine synthesis and catabolism.
Genes of Ti Plasmid and their Function
| vir | DNA transfer into plants |
| shi | Shoot induction |
| roi | Root induction |
| nos | Nopaline synthase |
| noc | Nopaline catabolism |
| ocs | Octopine synthase |
| occ | Octopine catabolism |
| tra | Bcterial transfer genes |
| inc | Incompatibility genes |
| oriV | Origin of replication |
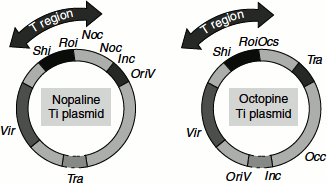
The Ti plasmids can be grouped into three types based on the opine genes they possess; for example, octapine, nopaline and agropine (Figure 9.23).
Figure 9.23 Ti plasmid
A. tumifaciens attacks many dicotyledonous plants and results in the formation of a tumour. The lipopolysaccharide secreted by the bacterial cell wall helps in its attachment with the polygalacturonic fraction of the plant cell wall. From the wounded cell wall of plant, a phenolic compound called acetosyringone is secreted, which induces the vir genes of Ti plasmid. Vir genes encode an enzyme that nicks the T-DNA and mediates its transfer into the plant cell and gets integrated with the plant DNA. Ops genes encodes enzymes for the synthesis of opines in the transformed cell, which are required for bacterial proliferation. The T-DNA also encodes enzymes that are involved in the biosynthesis of phytohormones, auxin and cytokinin. This results in the disorganized proliferation of plant cells that are commonly known as callus or galls or tumour. The galls are colonized by A. tumefaciens.
The Ti plasmid can act as a very good plant vector. The DNA of interest could be spliced into the T-DNA. The Ti plasmid cannot easily be manipulated. An intermediate vector initially receives the DNA of interest and various other genes necessary for recombination, replication and antibiotic resistance. The intermediate vector can then be inserted into the Ti plasmid forming a, co-integrated plasmid.
The Ti plaarmed, i.e., its entire right-hand region of its T-DNA, including tumour genes and nopaline synthase genes are deleted, making it in capable of tumour formation. It retains the left-hand border of its T-DNA, which is used as the cross-over site for the incorporation of the intermediate vector. The intermediate vector contains multiple unique restriction sites. The gene of interest can be introduced in these sites. The intermediate vector also carries suitable antibiotic marker genes such as kanamycin-resistant gene (kanR) for the selection of the recombinants.