The immediate products of transcription, the primary transcripts, are not necessarily biologically active. In order to make them as functional entities, many of them must be altered in several ways. These modifications are referred to as post-transcriptional modifications or processing, which include:
- Exo- and endonucleolytic removal of polynucleotide segments,
- Appending nucleotide sequence to their 3′-and 5′-ends and
- The modification of specific nucleosides.
mRNA Processing
Prokaryotic mRNA
In prokaryotes, most primary mRNA transcript functions in translation without further modification. Prokaryotes usually commence translation on nascent mRNAs themselves. Therefore, the prokaryotic mRNA does not undergo post-transcriptional processing.
Eukaryotic mRNA processing
In eukaryotes, mRNAs are synthesized in nucleus while translation occurs in the cytosol. Apart from spatial segregation, there also exists a finite temporal lag between transcription and translation (Figure 4.21).
Figure 4.21 mRNA transcription in the nucleus and its translation in the cytosol
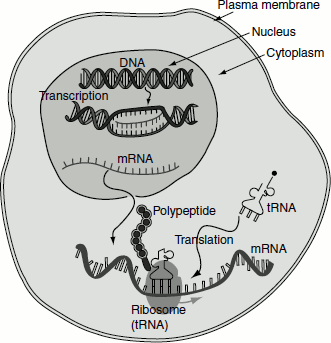
The mRNA is synthesized as heterogeneous nuclear RNA [hnRNA] or pre-mRNA. This is the primary mRNA transcript in the nucleus, which undergoes extensive post-transcriptional processing while still in the nucleus, to form mature mRNA and is then transported to cytosol to get associated with ribosomes for the translation process to commence. hnRNPs (ribonucleoproteins) associate with the primary transcript and prevent them from forming secondary structure and thus maintain it in a linear form, so that it undergoes translation.
Processing of mRNA involves the following events:
- Capping,
- Tailing,
- Splicing and
- Methylation.
Capping
All eukaryotic mRNAs have a cap structure at the 5′-end consisting of a 7-methylguanosine residue that is attached to the transcript via 5′-5′-triphosphates bridge by guanylyl transferase. The cap structure is attached to the 5′-end of the growing transcript before it is approximately 30 nucleotides long.
Types of capping
There are three types of capping.
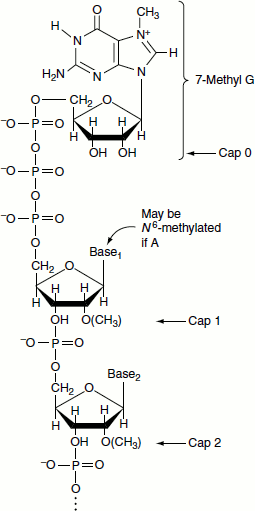
- Cap 1: In addition to the 7-methylguanosine cap, if the first nucleoside is methylated at the 2′ position, it is called cap 1 structure. It is the predominant capping in most of the multicellular organisms.
- Cap 2: When the first two nucleosides following 7-methylguanosine is methylated at 2′ position, it is called cap 2 structure and it is found in some eukaryotes.
- Cap 0: When the two leading nucleosides of the RNA transcript are not methylated at 2′ position and just bears the 7-methylguanosine cap, then it is referred to cap 0. It occurs predominantly in unicellular eukaryotes (Figure 4.22).
Figure 4.22 Reactions of mRNA capping
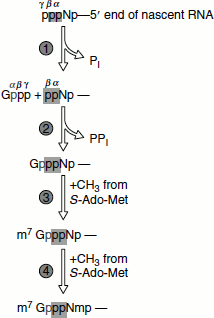
Capping involves several enzymatic reactions:
- Removal of the leading phosphate group from the mRNA’s 5′-terminal triphosphate group by an RNA triphosphatase.
- Guanylation of the mRNA by capping enzyme requires GTP and yields a 5′-5′-triphosphate bridge.
- Methylation of guanine by 7-methyltransferase, methyl group supplied by S-adenosyl methionine.
- 0-2′-methylation of the mRNA’s first and second nucleotides by 2′-O-methyltransferase (Figure 4.23).
Both capping enzyme and guanine-7-methyltransferase bind to RNA Pol-II’s phosphorylated CTD. Capping marks the completion of RNA Pol-II’s switch from transcription initiation to elongation.
Significance of capping
- Capping enhances the translation ability of mRNA. The capping of mRNA is believed to be essential for binding to the smaller subunit of ribosomes.
- The capping protects mRNA from ribonuclease (RNase).
Figure 4.23 Cap structure
Tailing
Eukaryotic mRNAs have poly(A) tails. The primary transcript of a given structural gene has heterogeneous 3′-sequences but mature mRNA has well-defined 3′-ends. Almost all of them have 3′-poly(A) tails. Tailing is a process in which poly(A) tail with around 200 adenosine residues is attached to 3′-end of hnRNA.
The poly(A) tails are enzymatically appended to the primary transcript in two reactions.
- Primary transcript is cleaved to yield free 3 -OH group at a specific site, 15–25 nucleotides past an AAUAAA sequence and within 50 nucleotides before a U-rich or G+U-rich sequence. The AAUAAA sequence is highly conserved. The mutations of this sequence abolish cleavage and polyadenylation. The precision of the cleavage reactions apparently eliminates the need for accurate transcriptional termination. The cleavage factors I and II are required for the process.
- Poly(A) tail is subsequently generated from ATP, by the action of Poly(A) polymerase (PAP). The enzyme is recruited by cleavage and poly adenylation specificity factor (CPSF). Downstream G+U-rich element is recognized by cleavage stimulation factor (CstF), which increases the affinity with which CPSF binds to the AAUAAA sequence.
Even in tailing are shown in Figure 4.24
Significance of polyadenylation
Experimental evidences are shown that poly(A) stabilizes mRNA. mRNAs that have poly(A) tail have greater lifetime in cytosol, whereas other mRNAs that have no poly(A) tail have lifetime less than 30 minutes in cytosol.
Splicing
The most striking differences between eukaryotic and prokaryotic structural genes are that the coding sequences of most eukaryotic genes are interspersed with unexpressed regions. Because of this, the eukaryotic genes known as ‘split genes’.
Splicing reaction involves the removal of introns and ligation of exons.
Exons: They are the coding or functional sequences or the expressed sequences of gene, which are transcribed in the primary RNA transcript and are retained in the final mature mRNA.
Introns: They are the non-coding or non-functional intervening sequences (IVs) of gene that are transcribed in the primary RNA transcript but are not retained in the mature mRNA because of splicing reactions.
Figure 4.24 mRNA tailing by PAP
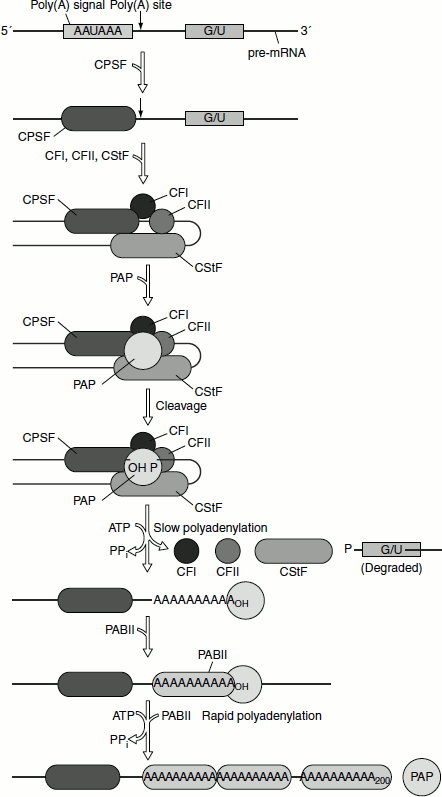
A typical mammalian gene has 7–8 exons spread out over approximately 16 Kb. The exons are relatively short (100-200 bp), while the introns are relatively long (>1 Kb). In general the splicing reactions are of three types (Figure 4.33).
Mechanism of splicing
Splice sites/splice junctions
These are sequences immediately surrounding the exon-intron boundaries. At the splice junction, ‘AAGU’ is the highly conserved sequence at the 5′-boundary (5′-splice site) and ‘AGG’ at the 3′-boundary (3′-splice site). In the introns, a conserved sequence of ‘CURAY’ [where R represent purines and Y represent pyrimidines], which is highly conserved in vertebrate mRNA, has been found about 20–50 residues upstream at the 3′-splice site. This sequence is called the ‘branch site’.
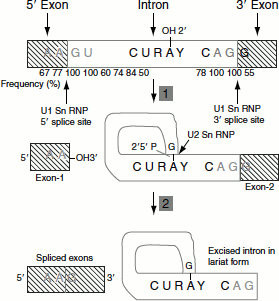
Steps in mRNA splicing
- A cut is made at the 5′-splice site.
- Left exon and right intron molecules separate.
- The left exon takes the form of a linear molecule.
- The right intron-exon molecule forms a lariat in which the 5′-terminus generated at the end of the intron, guanosine’s 5′-terminal phosphate, forms a 2′-5′-phosphodiester bond with the introns adenosine residue at the branch site.
- Now, free 3′-OH group of the 5′-exon forms a phosphodiester bond with the 5′-terminal phosphate of the 3′-exon, yielding the splice product. The introns are eliminated in its lariat form. The lariat intron is debranched to give linear excised intron, which is rapidly degraded (Figure 4.25).
Figure 4.25 Steps in mRNA splicing
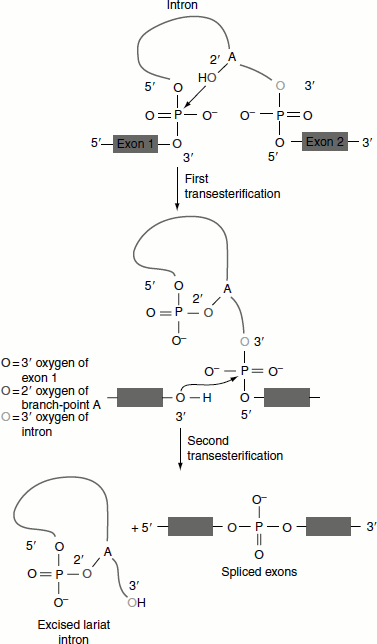
Chemistry of splicing reaction
Nuclear splicing occurs by two transesterification reactions in which an OH group attacks a phosphodiester bond.
- Nucleophilic attack by the 2′-OH of A of the branch site on the 5′-splice site.
- The free 3′-OH of the exon, released by the first reaction attacks the bond at the 3′-splice site (Figure 4.26).
Spliceosome
The splicing components associate to form a large complex called spliceosome. The spliceosome recognizes the 5′-and 3′-splice sites and the branch sites. The complex assembles sequentially on the pre-mRNA. The splicing apparatus contains both proteins and RNAs. The RNAs take the form of small molecules that exist as ribonucleoproteins particles. Both the nucleus and the cytoplasm of eukaryotic cells contain many discrete small RNAs. The size of these RNAs ranges from 100–300 bp. These ribonucleoparticles found in the nucleus are called ‘small nuclear RNAs (snRNA)’, while those found in the cytoplasm are called ‘small cytoplasmic RNAs (scRNA)’; also called Snurps and Scyrs. There are also a class of small RNAs found in the nucleolus called snoRNAs, which are involved in the processing of ribosomal RNA.
The snRNPs involved in splicing are U1, U2, U5, U4 and U6. They are named according to the snRNAs that are present. Each snRNP contains a single snRNA and several proteins.
Figure 4.26 Transesterifications in splicing
Splicing factors
A variety of proteins, which are known as splicing factors and are extrinsic to spliceosome, also participate in splicing. Among them are ‘branch point-binding protein (BBP also known as splicing factor 1) and U2 snRNP auxiliary factor (U2 AF)’, co-operate to select the branch point intron. U2 AF binds to the polypyrim-idine tract upstream of the 3′-splice site, whereas BBP recognizes the nearby branch point sequence.
Splicing proceeds in the 5′ → 3′ direction and occurs co-transcriptionally. Thus, when a newly synthesized exon emerges from RNA Pol-II, it is bound by splicing factors that are also bound to RNA Pol-II highly phosphorylated CTD. This attaches the exon and its associated spliceosome to the CTD to ensure that splicing occurs when the next exon emerges from RNA Pol-II.
Spliceosomal assembly
The binding of U1 snRNP to the 5′-splice site is the first step in splicing. U1 snRNA base pairs with the 5′-splice site by means of a single-stranded region at its 5′-terminus, which usually includes a stretch of four to six bases that is complementary with the 5′-splice site. The first complex formed during splicing is the E (early pre-splicing complex), which contains U1 snRNP, the splicing factor U2 AF and the SR proteins (Ser-Arg-rich proteins). These connect the U2 AF to U1. The E complex is sometimes called the commitment complex because its formation identifies a pre-mRNA as a substrate for splicing.
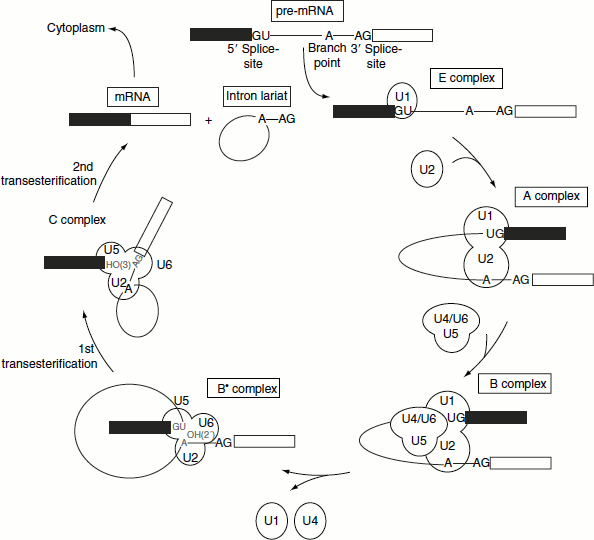
Following the formation of E complex, the other snRNPs and factors involved in splicing associate with the complex in a defined order. The B complex is formed when a trimer containing the U5, U4 and U6. snRNPs bind to the A complex containing U1, U2 and snRNPs. It is converted to the B∗ complex after U1 is released. The dissociation of U1 is necessary to allow other components to come into juxtaposition with the 5′-splice site, most notably U6 snRNP. Following this, U5 snRNP, which is initially close to exon sequences at the 5′-splice site, changes its position and shifts to the vicinity of the intron sequences, forming the C complex (Figure 4.27).
The catalytic reaction is triggered by the ATP hydrolysis-driven release of U4. The role of U4 snRNP may be to sequester U6 snRNP until it is needed. In the U6/U4 snRNP, a continuous length of 26 bases of U6 is paired with two separated regions of U4. When U4 dissociates, the region in U6 is released and becomes free, forming another structure. The first part of it pairs with U2. The second part forms an intramolecular hairpin. The base pairing between U2 and the branch point creates a catalytically active structure that carries out the process of splicing.
Figure 4.27 The spliceosomal assembly and splicing of pre-mRNA
Methylation
Approximately 0.1 per cent of the A residues of the pre-mRNAs are methylated at their N6 during or shortly after the synthesis.
RNA Editing
The sequence of a pre-mRNA is altered by a process called RNA editing. As a consequence the mature mRNA sequence differs from the exons encoding it in genomic DNA. These changes include C → U and U → C changes, the insertion or the deletion of U residues and the insertion of multiple G or C residues.
RNA editing occurs by two distinct mechanisms:
- Substitution editing: In this type of editing, the individual nucleotides are altered chemically (the equivalent of point mutations). Such changes are mediated by enzymes.For example, cytidine deaminases convert a C in the RNA to uracil (U) and adenosine deaminases convert an A to inosine (I), which the ribosome translates as a G. Thus, a CAG codon (for Gln) can be converted to a CGG codon (for Arg).
- Insertion/Deletion editing: This type of editing involves the insertion or deletion of nucleotides in the RNA. These alterations are mediated by guide RNA molecules that base pair as best they can with the RNA to be edited and serve as a template for the addition (or the removal) of nucleotides in the target.
RNA Editing of apoB mRNA
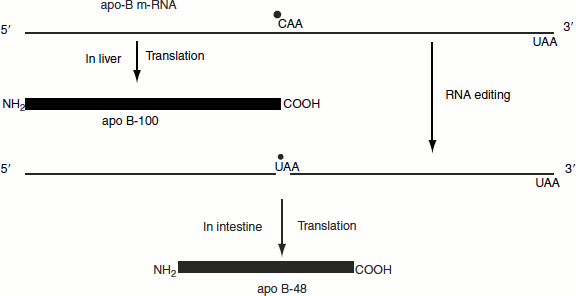
The intestine and liver have two different types of apoB lipoproteins produced by the same gene. The production of different apoBs from the same gene is because of RNA editing.
The apoB mRNA produced in the liver has the same sequences as the exons in the primary transcript that it is not RNA edited. This mRNA is translated into apoB-100.
The apoB mRNA produced in the intestine has the CAA codon in exon 26, which is edited to a UAA stop codon. As a result, intestine cells produce apoB-48. The addition of ‘U’ residues to mRNA is achieved with the help of guide RNAs (gRNA) during RNA editing (Figure 4.28).
Processing of rRNA
Processing of prokaryotic rRNA
The prokaryotic rRNAs are of three types, namely:
- 16S rRNA (1,541 ribonucleotides),
- 23S rRNA (2,904 ribonucleotides) and
- 5S rRNA (120 ribonucleotides).
Figure 4.28 RNA editing of apoB mRNA
The E. coli rRNA operons contain one copy each of the rRNA genes. Their polycistronic primary rRNA transcript is approximately 5,500 nucleotides long and contains 16S rRNA at their 5′-end, followed by the transcripts for one or two tRNAs, 23S rRNA, 5S rRNA and, in some RNA operons, one or more tRNAs at their 3′-end. The initial processing yields products known as pre-rRNAs.
Processing involves the following steps: (Figure 4.29)
- Primary processing,
- Secondary processing and
- Methylation.
Figure 4.29 Prokaryotic rRNA processing
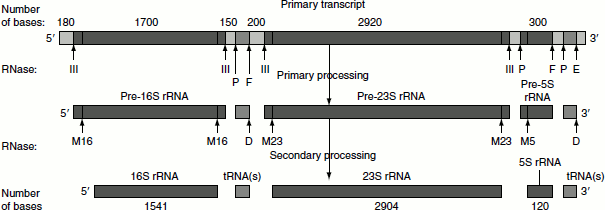
Primary processing
In this process, the primary rRNA undergoes endonucleolytic cleavages in which the rRNAs (16S, 23S and 5S) and tRNAs are cleaved by trimming of the flanking nucleotide sequences. The trimming is catalysed by RNase III, RNase P, RNase E and RNase F.
Secondary processing
The endonucleolytic activity of RNase III, P, E and F does not completely trim the flanking regions of the rRNAs. The 5′-and 3′-ends of 16S rRNA, 23S rRNA and 5S rRNA are further trimmed by the RNase M16, M23 and M5, respectively. RNase D involves in the trimming of flanking regions of tRNA. After secondary processing, the rRNAs get associated with proteins to form ribosomes.
Methylation
A total of 24 specific nucleotide residues of the 16S rRNA and 23S rRNA are methylated during the ribosomal assembly. The methylation reaction employs S-adenosyl methionine, a methyl donor, and yields N6, N6-dimethyladenine and 2′-O-methyl ribose residues.
These methylation reactions are thought to protect adjacent phosphodiester bond from the degradation by intracellular RNases.
Processing of eukaryotic rRNA
The eukaryotic genomes have several hundreds of tandemly repeated copies of rRNA genes that are contained in small and dark staining nuclear bodies known as nucleoli, which are the site of rRNA transcription, processing and ribosomal subunit assembly.
There are of three types of rRNA, namely:
- 18S rRNA (1,900 nucleotides),
- 5.8S rRNA (160 nucleotides),
- 28S rRNA (4,700 nucleotides).
The primary rRNA transcript is of approximately 7,500 nucleotide residues and has a sedimentation coefficient of 45S. Starting from 5′-end, the structural arrangement of various rRNAs in the pre-rRNA is as follows:
5′ — 18S rRNA —- 5.8S rRNA —- 28S rRNA —- 3′
As in prokaryotes, these rRNAs are separated by spacer sequences. The processing of these rRNAs involves the following steps and is guided by small nucleolar RNAs (snoRNAs). The processing can be divided into the following steps (Figure 4.30).
- Methylation,
- Primary processing,
- Secondary processing and
- Splicing.
Methylation
In the first stage of its processing, 45S rRNA is specifically methylated at numerous sites. About 80 per cent of these modifications yield O2′ methyl ribose residues and the remaining 20 per cent of modifications form methylated bases such as N6, N6-dimethyladenine and 2-methylguanine. In addition, the uracil residues of the rRNA are converted to pseudouridines. rRNAs interact with the members of small nucleolar RNAs (snoRNAs), which requires rRNA methylation.
Figure 4.30 Eukaryotic rRNA processing
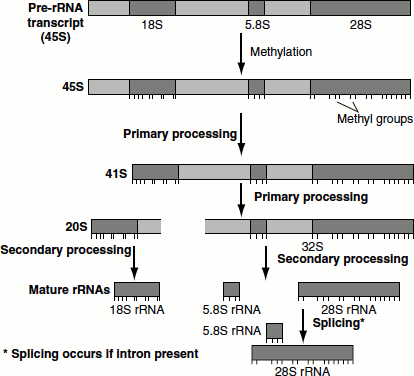
Primary processing
Soon after methylation, 45S rRNA undergoes cleavage at the 5′-end of the spacer to yield 41S rRNA. The 41S rRNA is further cleaved into two pieces, 32S and 20S, that contain the 28S and 18S sequences, respectively. The 32S precursor retains the 5.8S RNA sequence.
Secondary processing
32S precursor is split to yield the mature 28S and 5.8S RNAs, which base pair with each other and the 20S precursor is trimmed to mature 18S size.
Splicing
A few eukaryotic rRNA genes contain introns and undergo splicing reactions. For example, The 26S part of the protozoan Tetrahymena thermophilia’s rRNA precursor contains introns and it can be spliced by 26S rRNA itself without any help from proteins. The T. Thermophilia’s 26S rRNA is the equivalent of the mammalian 28S rRNA.
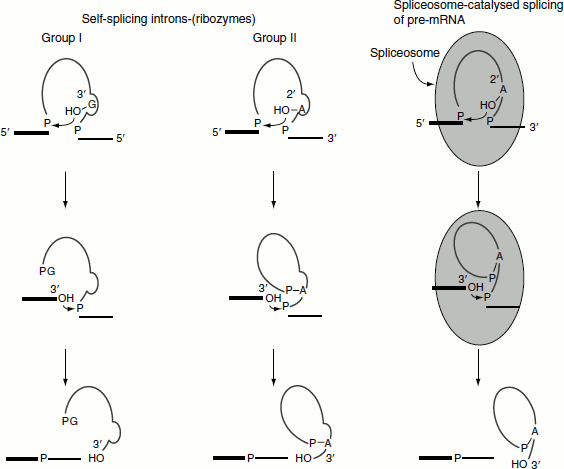
Group-I introns splicing
The introns in 26S rRNA of T. thermophilia are known as group-I introns, which also occur in the nuclei, mitochondria and chloroplast of diverse eukaryotes. The splicing of 26S rRNA was explained by Thomas Cech. The splicing reaction involves the following steps: (Figure 4.31)
- A guanine nucleotide attacks the adenine nucleotide residue at the 5′-end of the introns,
- Exon 1 is released from the rest of the molecule and leaves intron 1 and exon 2 complexes,
- Exon 1 attacks exon 2,
- Exon 1 and exon 2 are spliced and
- Linear intron is released.
Figure 4.31 Group-I introns splicing
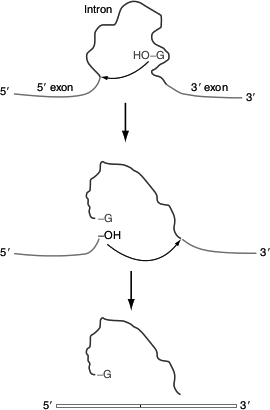
Figure 4.32 Group-II introns splicing
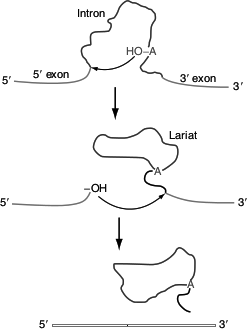
Group-II intron splicing
The introns of yeast mitochondrial pre-rRNA are known as group-II introns and are also found in the mitochondria of fungi and plants. They comprise the majority of the introns in chloroplasts. Group-II introns are also self-splice, but they do not need assistance from guanosine to start the reaction. Instead, the initiating molecule is an adenosine nucleotide residue within the introns of the RNA itself.
The splicing reaction involves the following steps: (Figure 4.32)
- 2′-OH of adenosine residue attacks the 5′-end nucleotide residue and forms a lariat structure with introns 1 and exon 2 complexes,
- Exon 1 is released,
- 3′-OH group of exon 1 attacks exon 2,
- Exons 1 and 2 are spliced and
- Lariat intron is released.
The details of the rRNA processing scheme are not universal. Still, the basic mechanism of rRNA processing, including the order of mature sequences in the precursor, is preserved throughout the eukaryotic kingdom.
Figure 4.33 Three Types of splicing reactions
Ribozymesw
RNAs with enzymatic activities are referred to as ribozymes; for example, hammerhead ribozymes of plant virus and T. thermophilia’s rRNA. Since splicing carried out by RNA itself, the process is known as self-splicing. Such RNA (group-I introns) enzymes are called ribozymes. This self-splicing process consists of a series of transesterifications and, therefore, does not require energy input. Other examples of ribozymes include the RNA component of RNase P.
For mechanism, refer to group-I intron splicing (Figures 4.31 and 4.33).
Processing of tRNA
Both prokaryotic and eukaryotic pre-tRNAs undergo post-transcriptional modification (Figure 4.34). The steps for the post-transcriptional modification of pre-tRNA are as follows:
- The 3′-OH and 5′-phosphate ends are cleaved by the endonuclease action of RNase D and RNase P, respectively.
- The introns in the anticodon loop are spliced.
- tRNA-specific nucleotidyl transferase adds the trinucleotide CCA is to the 3′-end.
- Base modifications to give mature tRNA.
Figure 4.34 tRNA processing
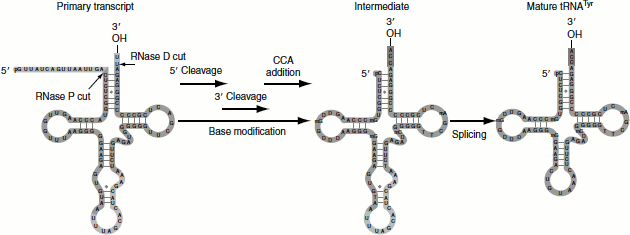
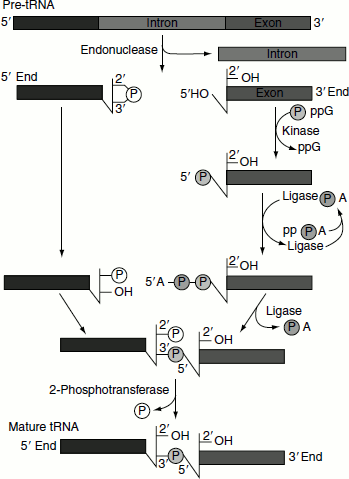
Figure 4.35 Mechanism of tRNA splicing
tRNA splicing
Splicing mechanism in pre-tRNA differs from the mechanisms utilized by self-splicing introns and spliceosomes.
tRNA splicing reaction is mediated by four enzymes; namely, tRNA-specific endonuclease, cyclic phosphodiesterase, tRNA-specific ligase and 2-phosphotransferase (Figure 4.35).
- An endonuclease removes the introns.
- 2′-3′-cyclic phosphomonoester bond forms on the cleaved end of the 5′-exon.
- 3′-exon is subjected to the action of kinase and ligase utilizing the energy provided by ATP and GTP hydrolyses.
- 5′-exon and 3′-exon are ligated.
- The 2′-phosphate on the 5′-exon is removed in the final step.