Binding of RNA Polymerase
RNA polymerase is sufficiently large enzyme that can come into contact with many DNA bases simultaneously. An estimate of the size of the region of the DNA where contact is made is obtained by selectively degrading adjacent DNA bases with DNase, a procedure known as ‘DNase protection method’.
Figure 4.7 DNA foot printing
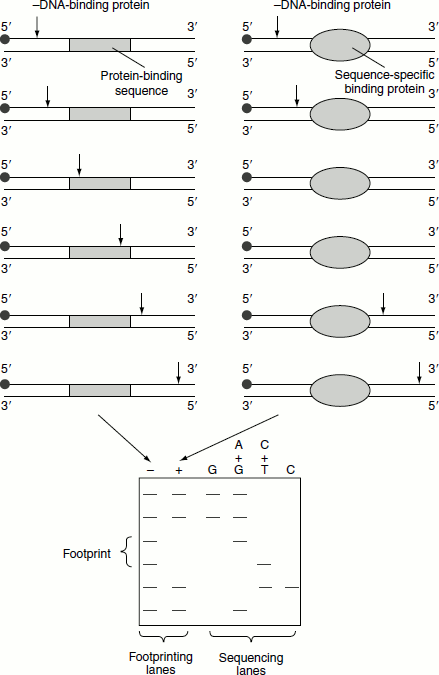
RNA polymerase is bound to DNA; then, a DNA endonuclease is added to the mixture. The nuclease degrades most of the DNA to mono- and dinucleotides but leaves untouched the DNA segments in close contact with the RNA polymerase. These segments vary from 41 to 44 bp.
DNA Foot Printing
‘Foot printing’ detects the regions of both strong binding and weak binding and shows that the complete contact region is about 70 bp. A particular piece of a double-stranded DNA having a known sequence of bp, binding RNA polymerase is labelled in one strand at its 5′-terminus with P32. It is then allowed to interact with the polymerase. An endonuclease is then added, but so briefly that no DNA molecule receives more than one single-strand break. The endonuclease nicks all regions except those protected by the RNA polymerase. The DNA is then isolated and denatured. The radioactively labelled DNA fragments will now consists of set of molecules that vary in length according to their distance from the 5′-end to the position of the endonuclease. If the DNA contains ‘n’ bp and the RNA polymerase is not added, the n size of the DNA fragments will be present. However, if the RNA polymerase binds to ‘x’ bp and thereby prevents the access of the DNA to the nuclease, only the n-x different size of the DNA fragments will be represented. These fragments are separated by gel electrophoresis. Upon comparing the DNA fragments of protein bound ones with that of the protein unbound DNA fragments, the protein bound DNA sequences can be identified (Figure 4.7).
Transcriptional Events
Transcription initiation
The first step in the transcription is the binding of RNA polymerase to a DNA molecule. Several factors in the sigma subunit and the CTD in the alpha subunit contact promoter DNA. The sigma factor contacts the promoter directly at both the −35 and −10 consensus sequences. The process can be broken into three parts:
- Template binding at a polymerase recognition site.
- Binding to an initiation site.
- Establishment of an open promoter complex (Figure 4.8).
Figure 4.8 Transcription initiation
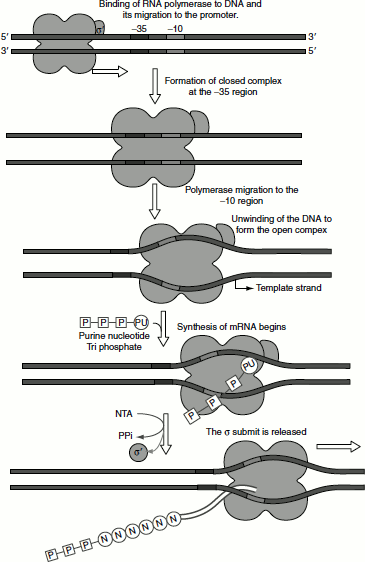
RNA polymerase binds to the promoter in at least two distinguishable steps. The holoenzyme first binds the DNA and migrates to the −35 region and forms a closed complex. The DNA is then unwound for about 17 bp starting from the −10 region. The unwinding exposes the template strand at the initiation site. The RNA polymerase binds more tightly to this unwound region, forming an open complex. The sigma subunit is required only to ensure the specific recognition of the promoter by the RNA polymerase. Once a few phosphodiester bonds are formed, the sigma subunit dissociates leaving the core polymerase to complete the synthesis of RNA molecule.
RNA polymerase contains two nucleotide-binding sites called the initiation site and the elongation site. The initiation site primarily binds purine triphosphates, ATP and GTP. ATP is usually the first nucleotide in the chain and is usually retained as a triphosphate. Thus, the first DNA base that is transcribed is usually thymine. The elongation site also called the catalytic site is then filled with nucleotide triphosphates that are selected by their ability to hydrogen bond with the next base of the DNA template. The two nucleotides are then joined together and the first base is released from the initiation site and initiation is complete.
The RNA polymerase and the template strand move relative to each other, so that the two binding sites and the catalytic site are shifted by exactly one nucleotide.
Chain elongation
After several nucleotides are added to the growing chain, RNA polymerase undergoes a conformational change and loses the sigma subunit. The core enzyme moves along the DNA extending the RNA chain. The DNA helix recloses, as the synthesis proceeds. The newly formed RNA is released from its hydrogen bonds with the DNA, as the helix reforms. Roughly, 12 RNA bases are paired to the DNA in the open region. The newly synthesized RNA forms a hybrid helix with the template DNA strand. The RNA-DNA helix is about 12 bp long. The 3′-OH of this helix is so positioned that it can attack the α-phosphate of the incoming NTP.
A peculiarity of the chain elongation is that it does not occur at constant rate, i.e., the synthesis markedly slows down when particular regions of DNA are passed, then continues at the normal rate, slows down again and accelerates again and so forth. This reduction in the transcription rate is called pause. Analysis of pausing along the stretches of the DNA of known sequences shows that the pausing frequently follows the sequences that form hairpins in the RNA. The pausing in the non-hairpin regions may be associated with an increase in the rate of phosphor ester cleavage by RNA polymerase. The RNA polymerase catalyses both polymerization and degrading with polymerization predominating vastly (Figure 4.9).
The elongation is about 50 nucleotides per second. The transcription bubble (the region containing RNA polymerase, DNA and nascent RNA) moves at a distance of 170 Å in this time. In contrast with DNA polymerase, RNA polymerase does not edit the nascent polynucleotide RNA chain. Consequently, the fidelity of transcription is much lower than that of replication.
Termination
Rho independent termination
In the termination phase, the formation of phosphodiester bond ceases, the RNA–DNA hybrid dissociates, the melted region of the DNA rewinds and the RNA polymerase releases the DNA. This event is precisely controlled. The transcribed regions of DNA templates contain stop signals (intrinsic terminator)—requires a G+C-rich region followed by an A-T-rich region. The RNA transcript of this DNA palindrome is self-complementary and hence its bases pair among themselves forming a hairpin stem and loop structure. This region is followed by a series of U residues. The RNA transcript ends within or just after this.
Figure 4.9 Transcription bubble-elongation of the RNA
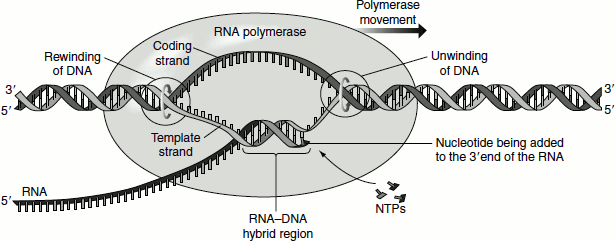
The RNA polymerase pauses when it encounters such hairpins. Furthermore, the RNA–DNA hybrid helix produced after the hairpin is unstable because of its content of rU–dA, which is the weakest of the four kinds of bp. Hence, RNA dissociates from DNA template and from the enzyme. The solitary DNA template strand rejoins its partner to form the duplex DNA. The core enzyme devoid of the sigma factor has less affinity for a single-stranded DNA and so leaves the DNA. Sigma rejoins the core enzyme to form the holoenzyme that can again search for a promoter to initiate a new transcript. This mode of transcription termination is called ‘intrinsic termination or rho-independent termination’.
Rho protein and termination
RNA polymerase needs to terminate transcription at a hairpin followed by several U residues without the help of other proteins. However, at other sites, termination requires the participation of an additional factor, called ‘rho protein’.
Rho hydrolyses ATP in the presence of a single-stranded RNA but not in the presence of a DNA or a duplex RNA. Rho is a hexamer of 46 kD that specifically bind a single-stranded RNA. A stretch of 72 nucleotides is bound by the rho protein, 12 per subunit. The rho is brought into action by the sequences present in a nascent RNA. The absence of simple consensus sequence indicates that the rho recognizes non-contiguous structural features. The ATPase activity of the rho enables it to move unidirectionally along the nascent RNA chain towards the transcription bubble. It then breaks the RNA-DNA helix by pulling RNA away (Figure 4.10). Proteins, in addition to rho, mediate termination; for example, nus A protein enables the RNA polymerase in E. coli, to recognize a characteristic class of termination sites.
Figure 4.10 A model for rho action on a nascent co-translated mRNA
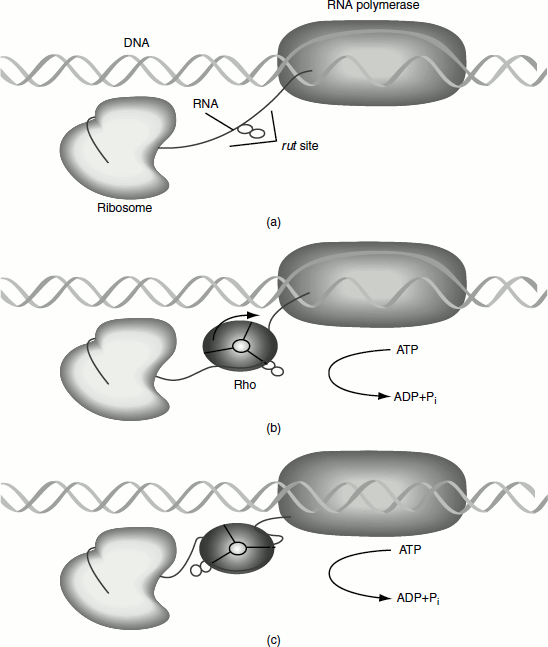
A common feature of rho-independent and -dependent terminations is the active signals lie in the newly synthesized RNA than in the DNA template.