All genes of the genome are not transcribed at all the time. In higher eukaryotes (such as human beings) especially, where there is a tissue-specific gene expression (such as muscle, for example), there is selective transcription of the genome. Therefore, it would be a tremendous waste of a cell’s resources to transcribe unnecessary genes. Transcription of genes needs to be controlled, so that they are transcribed only when and where they are needed. The regulatory regions of the DNA regulate the transcription of individual genes.
There are two major types of regulatory regions namely ‘promoters and enhancers’. Promoters are ‘molecular switches’ found immediately adjacent to the genes in the upstream direction. Transcription initiation takes place in the promoter region. Both prokaryotic and eukaryotic genes have promoter sequences. In prokaryotes, the sequence of a promoter is recognized by the ‘sigma (σ) factor’ of the RNA polymerase. In eukaryotes, it is recognized by specific ‘transcription factors’ (Table 4.3).
Table 4.3 E. coli σ factors and the consensus sequences of their recognition sites (promoters)
| σ Factor | Promoter consensus sequence | |
|---|---|---|
| -35 Region | -10 Region | |
| σ70 | TTGACA | TATAAT∗ |
| σ32 | TCTCNCCCTTGAA | CCCCATNTA |
| σ28 | CTAAA | CCGATAT |
| -24 Region | -12 Region | |
| σ54 | CTGGNA | TTGCA |
∗ 10 region is also called Pribnow box, after its discoverer.
† N = any.
Figure 4.6 Prokaryotic promoter
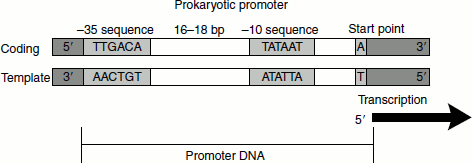
The regulatory protein interacts with the ideal sequences of the DNA called the ‘consensus sequence’, for example, the promoter sequence. These promoter consensus sequences are recognized by the RNA polymerase holoenzyme. A six-bp consensus promoter sequence located at the 10-bp upstream of the transcription start site is called the ‘−10 element’, ‘the Pribnow box’ or the ‘TATA box’. It has the consensus sequence TATAAT. The initially highly conserved TA and the final, almost completely conserved, T of the −10 element are crucial for the recognition of promoter. The conserved hexamer located at the −35-bp upstream of the start point is called the −35 element. The consensus is TTGACA. Bases in this sequence interact directly with the sigma factor. The distance separating the −35 and −10 sites is between 16 and 18 bp in approximately 90 per cent of the promoters (Figure 4.6). The 10–20 bp directly upstream of the −35 element is referred to as ‘UP element’. This region interacts with the (Carboxy Terminal Domain) CTDs of α-subunits of RNA polymerase.
Promoter Types
Promoters are classified as ‘high-level or strong promoters’. There are also ‘weak promoters’ in which the binding of RNA polymerase is less strong. In a given period of time, the number of RNA molecules synthesized from genes with weak promoters is much less than from a strong promoter. The promoter strength is one factor that determines the number of copies of each protein molecule present in the cell. The difference between strong and weak promoters lies in the sequence of bases at the −35 and −10 regions.