Physical Properties of the DNA
Size and shape
The DNA molecules range in length from about 2 microns (virus) to 2.1cm (Drosophila’s largest chromosome) to 1.6—8.2 cm (human chromosome) (1 kb DNA = 103 base pairs = 0.34 × 10-6 m). Some DNA molecules are circular (E. coli chromosome, mitochondrial DNA, plasmidDNA) and some are linear (human DNA, T7 DNA, lambda phage DNA). DNA molecules can be supercoiled or relaxed.
DNA denaturation and renaturation
The unwinding and separation of DNA strands is referred to as denaturation or ‘melting’. Denaturation can be induced experimentally. For example, if a solution of DNA is heated, the thermal energy increases molecular motion; this breaks the hydrogen bonds and other forces that stabilize the double helix and consequently the strands separate.
The denaturation of the double-stranded DNA can be followed spectroscopically. The purine and pyrimidine bases in the DNA absorb UV light maximally at a wavelength of approximately 260 nm. In the double-stranded DNA, this UV absorption is decreased due to base-stacking interactions. When a DNA is denatured, these interactions are disrupted and, as the result, an increase in absorbance is seen. This change is called the hyperchromic effect. The extent of the effect can be monitored as a function of temperature. Thus, as the DNA denatures its absorption of UV light increases. Near the denaturation temperature, a small increase in the temperature causes a simultaneous loss of the multiple, weak and co-operative interactions holding the two strands together, so that denaturation rapidly occurs throughout the entire length of the DNA (Figure 1.4).
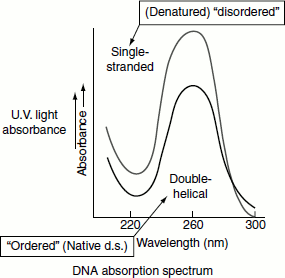
The temperature at which the strands of 50 per cent of a DNA molecule will separate is called the melting temperature, Tm. Tm depends on several factors. Molecules that contain a greater proportion of G≡C pairs require higher temperatures to denature because the three hydrogen bonds in G≡C pairs make them more stable than A═T pairs with two hydrogen bonds. The percentage of G≡C base pairs in a DNA sample can be estimated from its Tm (Figure 1.4). In addition to heat, the solutions of low ion concentration also favour DNA denaturation, causing it to melt at lower temperatures. DNA is also denatured by exposure to other agents that destabilize hydrogen bonds, such as alkaline solutions and the concentrated solutions of formamide or urea.
Figure 1.4 DNA denaturation and renaturation and melting tempertaure of DNA
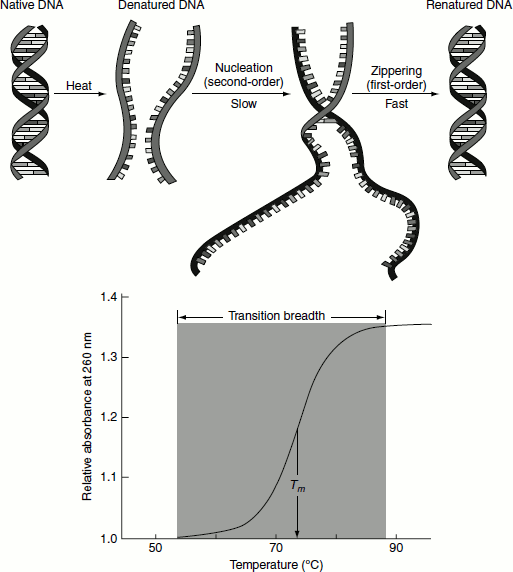
The single-stranded DNA molecules that result from denaturation form random coils without a regular structure. Lowering the temperature or increasing the ion concentration causes the two complementary strands to re-associate to form the double helix. The extent of such ‘renaturation’ is dependent on time, the DNA concentration and the ionic content of the solution. Two DNA strands that are not complementary in sequence will remain as random coils and will not renature. Further, they will not greatly inhibit complementary DNA partner strands from finding each other. The renaturation of melted DNA results in the decrease in UV absorption. This phenomenon is called ‘hypochromicity’.
The principles of DNA denaturation and renaturation form the basis of nucleic acid hybridization. The nucleic acid hybridization technique is used to study the relatedness of two DNA samples and to detect and isolate specific DNA molecules in a mixture containing numerous different DNA sequences.
Chemical Stability of Nucleic Acids
Hydrolysis by acids and alkali
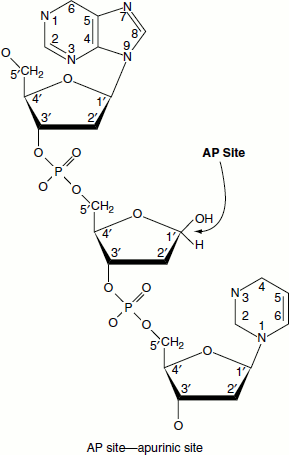
DNA is generally quite stable and resists the action of acid and alkaline solutions. However, in mild acid solutions—at pH 4—the beta-glycosidic bonds to the purine bases are hydrolysed. Protonation of purine bases (N7 of guanine and N3 of adenine) occurs at this pH. The protonated purines undergo hydrolysis. Once depurinated, the sugar can be easily isomerized into the open-chain form and in this form the depurinated (or apurinic) DNA is susceptible to cleavage. DNA is thus acid labile.
In contrast to DNA, RNA is very unstable in alkali solutions due to the hydrolysis of the phophodiester backbone. The 2′-OH group in ribo-nucleotides renders RNA molecules susceptible to strand cleavage in alkali solutions. Thus, RNA is alkali labile.
The alkaline hydrolysis of RNA results in the equimolar mixture of 2′- and 3′-nucleoside mono-phosphates.
Hydrolysis by enzymes
Enzymatic hydrolysis of RNA: There are many enzymes that cleave RNA—‘ribonucleases’.
Enzymatic hydrolysis of DNA: DNA is hydrolysed by ‘deoxyribonucleases’. These enzymes may digest a DNA strand from the end(s)—exonucleases—or internally—endonucleases (Table 1.4).
Salt concentration
The negatively charged phosphate groups in the DNA double helix are close together and will tend to repel one another unless they are neutralized. Since the concentration of salt (cations) in solution will affect the degree of neutralization, the stability of the DNA double helices also depends on the salt concentration. Salt ions, polyamines and special DNA-binding proteins help in the neutralization, which is taking place inside the cell.
Table 1.4 Nucleases
| Exonucleases | ||
| 5′ → 3′ exonucleases | Exonuclease VII, Bal31 nuclease | |
| 3′ → 5′ exonucleases | DNA polymerase I, Exonuclease l | |
| Endonucleases | ||
| Non-specific endonucleases | DNase I, Micrococcal nuclease, Mung bean nuclease | |
| Specific endonucleases | Restriction enzymes | |
pH
At a very high pH, the DNA will denature as single-stranded DNA. This is a commonly employed technique and is called alkaline gel electrophoresis. Low pH depurinates DNA (see hydrolysis by acids and alkali), which denatures the double helix. At very low pH, the phosophodiester back-bone of DNA hydrolyses forming nucleotides and nucleosides.
Ionic interactions
Proteins interact with DNA through ionic interactions. For example, the proteins called histones interact with the DNA. Arg and Lys can bind to any of the bases, other than C, by H-bond. However, these two residues almost always bind to G even through a single contact. This is probably due to ionic effects on the contacts. The G base has two acceptors and a partial negative charge, while T base is less negative, as it has only one acceptor. The A base has an acceptor and a donor and is nearly neutral; therefore, Arg and Lys bind to G much more often than to T, and to T more often than to A. His is less charged than Arg and Lys and thus its binding preference is expected to be weaker. The DNA charges may be neutralized by polypeptide chains that are bound to the grooves of the DNA double helix.
Mutagenic capacity of the bases
In general, the bases are stable because they are sequestered inside the double helix. However, notably two reactions can occur.
- Oxidative deamination of amino groupse.g. Cytosine → uracil
Adenine → hypoxanthine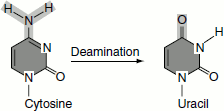
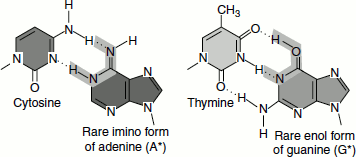
- Tautomerization: Sometimes the bases tautomerize to less common forms, (ex.) imino form of adenine, enol form of guanine.
Both reactions (i) and (ii) can affect the base-pairing potential of the bases.