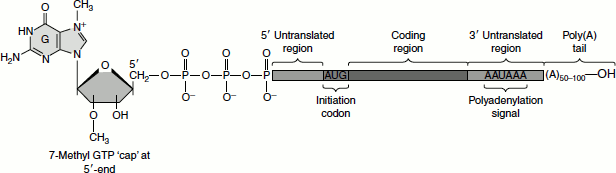
Eukaryotic mRNAs are characterized by two post-transcriptional modifications: the 5′-7-methyl-GTP cap and the poly(A) tail. The 7-methyl-GTP cap is essential for ribosomal binding of mRNAs in eukaryotes and also enhances the stability of these mRNAs by preventing their degradation by 5′-exonucleases (Figure 5.14). The poly(A) tail enhances both the stability and translational efficiency of eukaryotic mRNAs. The Shine-Dalgarno sequences found at the 5′-end of prokaryotic mRNAs are absent in eukaryotic mRNAs.
Figure 5.14 Capped and tailed eukaryotic mRNA
Initiation of Translation in Eukaryotes
The eukaryotic initiator tRNA is a unique tRNA functioning only in initiation. Like the prokaryotic initiator tRNA, the eukaryotic version carries only Met. However, unlike prokaryotic f-Met-tRNAfMet, the Met on this tRNA is not formylated. Hence, the eukaryotic initiator tRNA is usually designated tRNAiMet, with the ‘i’ indicating ‘initiation.’
Eukaryotic initiation can be divided into four fundamental steps.
- A ribosome must dissociate into its 40S and 60S subunits.
- A ternary complex termed the preinitiation complex is formed consisting of the initiator, GTP, eIF-2 and the 40S subunit.
- The mRNA is bound to the preinitiation complex.
- The 60S subunit associates with the preinitiation complex to form the 80S initiation complex.
The initiation factors eIF-1 and eIF-3 bind to the 40S ribosomal subunit favouring anti-association to the 60S subunit. The prevention of subunit re-association allows the preinitiation complex to form.
The first step in the formation of the preinitiation complex involves the binding of GTP to eIF-2 to form a binary complex. eIF-2 is composed of three subunits— α, β and γ. The binary complex then binds to the activated initiator tRNA; met-tRNAmet forming a ternary complex that then binds to the 40S subunit forming the 43S preinitiation complex. The preinitiation complex is stabilized by the earlier association of eIF-3 and eIF-1 to the 40S subunit (Table 5.2).
The cap structure of eukaryotic mRNAs is bound by specific eIFs prior to association with the preinitiation complex. Cap binding is accomplished by the initiation factor eIF-4F. This factor is actually a complex of three proteins—eIF-4E, A and G. The protein eIF-4E is a 24-kDa protein which physically recognizes and binds to the cap structure. eIF-4A is a 46-kDa protein which binds and hydrolyses ATP and exhibits RNA helicase activity. Unwinding of mRNA secondary structure is necessary to allow access of the ribosomal subunits. eIF-4G aids in binding of the mRNA to the 43 S preinitiation complex.
Once the mRNA is properly aligned onto the preinitiation complex and the initiator met-tRNAmet is bound to the initiator AUG codon (a process facilitated by eIF-1) the 60S subunit associates with the complex. The association of the 60S subunit requires the activity of eIF-5 which has first bound to the preinitiation complex. The energy needed to stimulate the formation of the 80S initiation complex comes from the hydrolysis of the GTP bound to eIF-2. The GDP-bound form of eIF-2 then binds to eIF-2B which stimulates the exchange of GTP for GDP on eIF-2. When GTP is exchanged, eIF-2B dissociates from eIF-2. This is termed as the eIF-2 cycle (see Figure 5.15). This cycle is absolutely required in order for eukaryotic translational initiation to occur. The GTP exchange reaction can be affected by phosphorylation of the α-subunit of eIF-2.
Table 5.2 Eukaryotic initiation factors and their functions
| Initiation factor | Activity |
|---|---|
| eIF-1 | Repositioning of met-tRNA to facilitate mRNA binding |
| eIF-2 | Ternary complex formation |
| eIF-2A | AUG-dependent met-tRNAmeti binding to 40S ribosome |
| eIF-2B (also called GEF) | GTP/GDP exchange during eIF-2 recycling |
| eIF-3, composed of 13 subunits | Ribosome subunit anti-association by binding to 40S subunit; eIF-3e and eIF-3i subunits transform normal cells when overexpressed; eIF-3A (also called eIF3 p170) overexpression has been shown to be associated with several human cancers |
| Initiation factor complex often referred to as eIF-4F composed of three primary subunits: eIF-4E, eIF-4A and eIF-4G and at least two additional factors: PABP and Mnk1 (or Mnk2) | mRNA binding to 40S subunit, ATPase-dependent RNA helicase activity, interaction between poly(A) tail and cap structure |
| PABP: poly(A)-binding protein | Binds to the poly(A) tail of mRNAs and provides a link to eIF-4G |
| Mnk1 and Mnk2 eIF-4E kinases | Phosphorylate eIF-4E increasing association with cap structure |
| eIF-4A | ATPase-dependent RNA helicase |
| eIF-4E | 5′-cap recognition; frequently found overexpressed in human cancers, inhibition of eIF4E is currently a target for anti-cancer therapies |
| 4E-BP (also called PHAS) three known forms | When de-phosphorylated, 4E-BP binds eIF-4E and represses its activity, phosphorylation of 4E-BP occurs in response to many growth stimuli leading to the release of eIF-4E and increased translational initiation |
| eIF-4G | Acts as a scaffold for the assembly of eIF-4E and eIF-4A in the eIF-4F complex, interaction with PABP allows 5′-end and 3′-ends of mRNAs to interact |
| eIF-4B | Stimulates helicase, binds simultaneously with eIF-4F |
| eIF-5 | Release of eIF-2 and eIF-3, ribosome-dependent GTPase |
| eIF-6 | Ribosome subunit anti-association |
Figure 5.15 Eukaryotic translation initiation
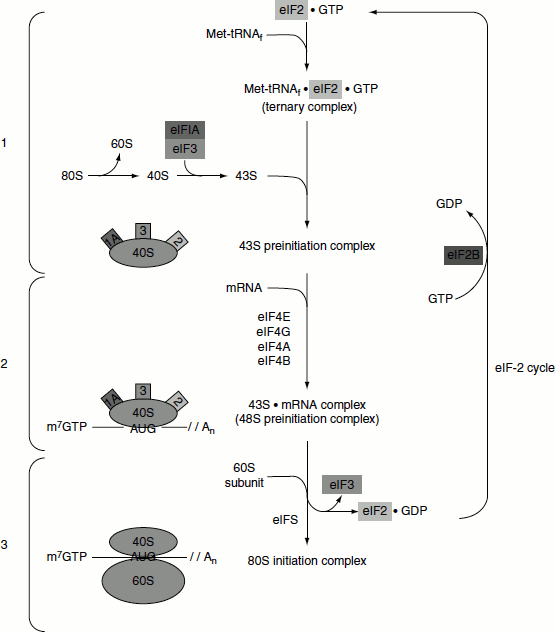
The eIF-2 cycle involves the regeneration of GTP-bound eIF-2 following the hydrolysis of GTP during translational initiation. When the 40S preinitiation complex is engaged with the 60S ribosome to form the 80S initiation complex, the GTP bound to eIF-2 is hydrolysed providing energy for the process. In order for additional rounds of translational initiation to occur, the GDP bound to eIF-2 must be exchanged for GTP. This is the function of eIF-2B which is also called GEF.
At this stage, the initiator met-tRNAmet is bound to the mRNA within the ribosome P-site. The incoming charged tRNAs binds to the A-site.
Elongation
The process of elongation, like that of initiation, requires specific non-ribosomal proteins namely EFs in prokaryotes; moreover, in eukaryotes, these are eEFs. Elongation of polypeptides occurs in a cyclic manner such that at the end of one complete round of amino acid addition, the A-site will be empty and ready to accept the incoming aminoacyl-tRNA dictated by the next codon of the mRNA. The process is accompanied by the movement of the ribosome to the next codon in the mRNA. Each incoming aminoacyl-tRNA is brought to the ribosome by an eEF-1 α-GTP complex. After the correct tRNA is deposited into the A-site, the GTP is hydrolysed and the eEF-1α-GDP complex dissociates. The GDP must be exchanged for GTP for additional translocation events. This is carried out by eEF-1βγ similarly to the GTP exchange that occurs with eIF-2 catalysed by eIF-2B.
The peptide attached to the tRNA in the P-site is transferred to the amino group at the amino-acyl-tRNA in the A-site. This reaction is catalysed by peptidyl transferase. This process is termed transpeptidation. The elongated peptide now resides on a tRNA in the A-site. The A-site needs to be freed in order to accept the next aminoacyl-tRNA. The process of moving the peptidyl-tRNA from the A-site to the P-site is termed as translocation. Translocation is catalysed by eEF-2 coupled to GTP hydrolysis. In the process of translocation, the ribosome is moved along the mRNA such that the next codon of the mRNA resides under the A-site. Following translocation, eEF-2 is released from the ribosome. The cycle can now begin again.
The ability of eEF-2 to carry out translocation is regulated by the state of phosphorylation of the enzyme, when phosphorylated the enzyme is inhibited. Phosphorylation of eEF-2 is catalysed by the enzyme eEF2 kinase (eEF2K). Regulation of eEF2K activity is normally under the control of insulin and Ca2+ fluxes. The Ca2+-mediated effects are the result of calmodulin interaction with eEF2K. Activation of eEF2K in skeletal muscle by Ca2+ is important to reduce consumption of ATP in the process of protein synthesis during periods of exertion which will lead to release of intracellular Ca2+ stores. eEF2K itself is also regulated by phosphorylation and one of the kinases that phosphorylates the enzyme is regulated by mTOR In addition, the master metabolic regulatory kinase, AMP-activated protein kinase, will phosphorylate and activate eEF2K leading to inhibition of eEF-2 activity.
Termination
Translational termination requires specific protein factors identified as releasing factors, known as eRFs in eukaryotes. There are two RFs in E. coli and one in eukaryotes. The termination signals are the same in both prokaryotes and eukaryotes. There are three termination codons, which are UAG, UAA and UGA.
The eRF binds to the A-site of the ribosome in conjunction with GTP. The binding of eRF to the ribosome stimulates the peptidyl transferase activity to transfer the peptidyl group to water instead of an aminoacyl-tRNA. The resulting uncharged tRNA left in the P-site is expelled with concomitant hydrolysis of GTP. The inactive ribosome then releases its mRNA and the 80S complex dissociates into the 40S and 60S subunits, ready for another round of translation.
Regulation of Translation
The expression of most genes is controlled at the level of their transcription. Transcription factors (proteins) bind to promoters and enhancers turning on (or off) the genes they control.
However, gene expression can also be controlled at the level of translation.
By general RNA-degradation machinery
P-Bodies
The cytosol of eukaryotes contains protein complexes that compete with ribosomes for access to mRNAs. As these increase their activity, they sequester mRNAs in larger aggregates called ‘P-bodies’.
These protein complexes break down the mRNA by:
- removing its ‘cap’,
- removing its poly(A) tail and
- degrading the remaining message (Degrading in the 5′ → 3′ direction).
What controls the dynamic balance between ribosomes and P-bodies for access to mRNAs remains to be learned. However, this mechanism provides for destruction of ‘bad’ mRNAs (e.g., those with premature STOP codons thus increasing the flexibility of gene expression in the cell.
Exosomes
These are hollow macromolecular complexes with two openings. They take in unfolded RNA molecules and degrade them in the 3′ → 5′ direction.
By MicroRNAs (miRNAs)
Here, small RNA molecules bind to a complementary portion in the 3′-UTR of the mRNA and prevent it from being translated by ribosomes and/or trigger its destruction. Both these activities take place in P-bodies.
By Riboswitches
The regulation of the level of certain metabolites is controlled by riboswitches. A riboswitch is a part of a molecule of mRNA with a specific binding site for the metabolite (or a close relative).
Examples:
- If thiamine pyrophosphate (the active form of thiamine [vitamin B1]) is available in the culture medium of E. coli,
- it binds to an mRNA whose protein product is an enzyme that is needed to synthesize thiamine from the ingredients in minimal medium.
- binding induces an allosteric shift in the structure of the mRNA, so that it can no longer bind to a ribosome and thus cannot be translated into the enzyme.
- E. coli no longer wastes resources on synthesizing a vitamin that is available preformed.
By gene-specific proteins
Translation of mRNA in humans is repressed by aminoacyl-tRNA synthetase in response to the inflammatory cytokine interferon-gamma [IFN-γ]. In presence of IFN-γ, the synthetase abandons its normal function (adding Glu and Pro to their respective tRNAs) and instead binds to the mRNA blocking its translation.
In some bacteria, a protein product may inhibit the further translation of its own mRNA (a kind of feedback inhibition). It does so by binding to a site which blocks the mRNA from further association with a ribosome.