Protein biosynthesis in all cells is characterized by three distinct phases: initiation, elongation and termination. At each stage, the energy required for the process is provided by GTP hydrolysis. Specific soluble protein factors participate in the events.
Activation of the Amino Acids
Amino acid activation takes place in the cytosol and not on the ribosomes. Each of the 20 amino acids is covalently attached to a specific tRNA. ATP provides the energy required. These reactions are catalysed by a group of Mg2+-dependent activating enzymes called ‘aminoacyl-tRNA synthetases’, each specific for one amino acid and its corresponding tRNAs.
When two or more tRNAs exist for a given amino acid, one aminoacyl-tRNA synthetase generally aminoacylates all of them. Aminoacylated tRNAs are referred to as being ‘charged’. Only the L amino acids take part in protein synthesis. Thus, a second genetic code is constituted by the aminoacyl-tRNA synthetases. That is each aminoacyl-tRNA synthetase discriminates between the 20 amino acids and the many tRNAs and uniquely picks out its proper substrates—one specific amino acid and the tRNA(s) appropriate to it—from among the more than 400 possible combinations.
Many of the other common amino acids which are not used in protein synthesis, e.g. citrulline, β alanine, etc. are also rejected. The activation of amino acids takes place through their carboxyl groups.
Amino acid + tRNA + ATP + Mg2+ → aminoacyl-tRNA +AMP + PPi
The activation reaction occurs in two steps. In the first step, an enzyme-bound intermediate, aminoacyl-adenylate (aminoacyl-AMP) is formed by reaction of ATP and the amino acid. In this reaction, the carboxyl group of the amino acid is bound in anhydride linkage with the 5′-phosphate group of AMP, with the displacement of pyrophosphate. In the second step, the aminoacyl group is transferred from the enzyme-bound aminoacyl-AMP to its corresponding specific tRNA (Figure 5.6).
The Two Classes of Aminoacyl-tRNA Synthetases
Aminoacyl-tRNA synthetases are a diverse group of proteins in terms of size, amino acid sequence and oligomeric structure. The aminoacyl-tRNA synthetases are classified into two fundamental classes based on similar amino acid sequence motifs, oligomeric state and acylation function, namely: class I enzymes, which are chiefly monomeric and class II aminoacyl-tRNA synthetases, which are always oligomeric (usually homodimers). Furthermore, first, class I aminoacyl-tRNA synthetases add the amino acid to the 2′-OH of the terminal adenylate residue of tRNA before shifting it to the 3′-OH; then, class II enzymes add it directly to the 3′-OH (Figure 5.7).
Figure 5.6 Amino acylation of tRNA by amino acyl tRNA synthetases
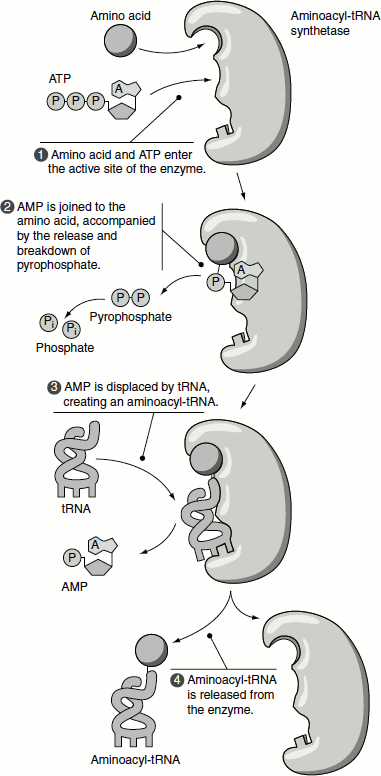
Figure 5.7 (a) t-RNA amino acylation (b) Steps in amino acylation by the two classes of amino acyl tRNA synthetases.
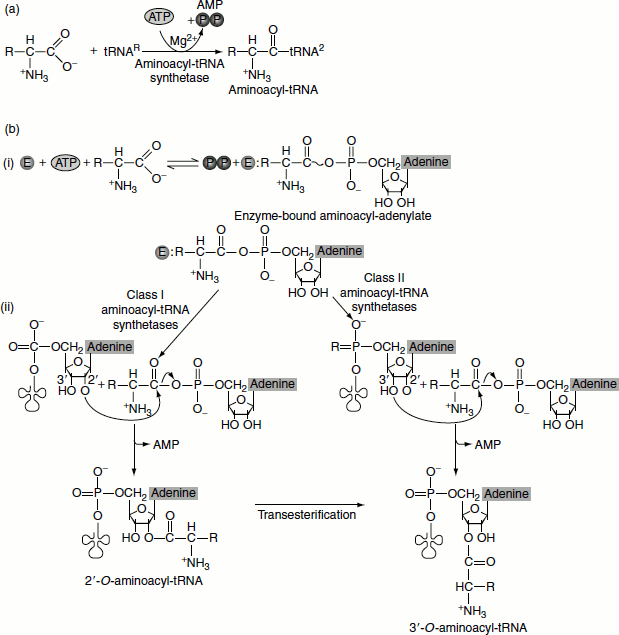
- activation of the amino acid.
- attachment of activated aminoacid to be tRNA
Aminoacyl-tRNA Synthetases and Proof Reading
While selecting an amino acid, aminoacyl-tRNA synthetases hardly make mistakes 1 in 10,000–100,000; however, while selecting tRNA, they may make a mistake 1 in 1,000,000. If a mistake has occurred, it is corrected by cognate tRNA binding; it also performs what is called chemical proof reading, i.e., after charging, if found wrong, it is hydrolysed and removed.
tRNA Molecule Acts as Adaptors
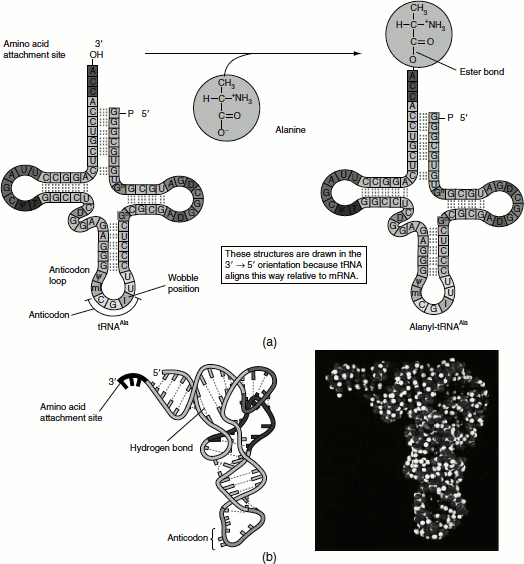
The fidelity of protein synthesis requires the accurate recognition of three-base codons on mRNA. An amino acid cannot itself recognize a codon. Consequently, an amino acid is attached to a specific tRNA molecule that can recognize the codon by Watson-Crick base pairing, that is tRNA serves as the adaptor molecule that binds to a specific codon and brings with it an amino acidfor incorporation into the polypeptide chain (Figure 5.8). Apart from tRNA molecules, the aminoacyl-tRNA synthetase enzymes are also adaptors of equal importance to the decoding process. Thus, the genetic code is translated by two sets of adaptors that act sequentially. Each of the adaptors match one molecular surface to another with great specificity and their combined action that associates, each codon with its particular amino acid.
Figure 5.8 Aminoacylated tRNA (a) Secondary structure of tRNA, before and after amino acid attachment (b) Tertiary structure of tRNA
The Interaction Between Aminoacyl-tRNA Synthetases and tRNA Constitutes a Second Genetic Code
An individual aminoacyl-tRNA synthetase must be specific not only for a single amino acid but for tRNA as well. Discriminating among several tRNAs is important for the overall fidelity of protein biosynthesis. The interaction between amino acyl-tRNA synthetases and tRNA has been referred to as ‘second genetic code’ to reflect its critical role in maintaining the accuracy of protein synthesis.
Polypeptide Synthesis Begins at the Amino-Terminal End
Polypeptide synthesis begins at the amino-terminal end and is elongated by sequential addition of residues to the carboxy-terminal end. This pattern has been confirmed by numerous researches and applies to all proteins in all cells.
The Process of Translation
Initiation
Initiation involves the reactions that precede the formation of the peptide bond between the first two amino acids of the protein. It requires the ribosome to bind to the mRNA. This is relatively slow step in translation and usually determines the rate at which an mRNA is translated. Initiation of translation is not a function of intact ribosomes, but is undertaken by the separate subunits, which re-associate during the initiation reaction.
The initiation codon in an mRNA is AUG, which codes for the amino acid methionine. There are two tRNAs for methionine in all organisms. One tRNA is used exclusively when AUG represents the initiation codon for protein synthesis. The second is used when methionine is added at an internal position in a polypeptide. In bacteria, two separate classes of tRNA specific for methionine are designated as tRNAMet and tRNAfMet. The starting amino acid at the amino-terminal end is N-formyl methionine. It enters the ribosome as N-formyl methionyl-tRNAfMet, which is formed in two successive reactions catalysed by the enzyme Met-tRNA synthetase.
Methionine + tRNAfMet + ATP → Met-tRNAfMet + AMP + PPi
Second, a formyl group is transferred to the amino group of methionine residue from N10-formyl tetrahydrofolate by a transformylase enzyme.
N10-formyl tetrahydrofolate + Met-tRNAfMet → tetrahydrofolate + fMet-tRNAfMet
This transformylase enzyme is more selective than the Met-tRNA synthetase and it cannot formylate free methionine residues or methionine residues attached to tRNAMet. Instead, it is specific for tRNAfMet.
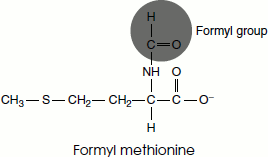
The initiation of polypeptide synthesis in bacteria requires:
- the 30S ribosomal subunit, which contains the 16S rRNA,
- the mRNA coding for the polypeptide to be made,
- the initiating fMet-tRNAfMet,
- a set of three proteins called initiation factors (IF-1, IF-2 and IF-3),
- GTP,
- the 50S ribosomal subunit and
- Mg2+.
The formation of the initiation complex takes place in three steps:
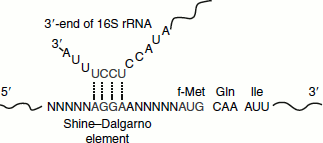
- The 30S ribosomal subunit binds initiation factor 3(IF-3), which prevents the 30S and 50S subunits from combining prematurely. Binding of the mRNA to the 30S subunit then takes place in such a way that the initiation codon AUG binds to a precise location on the 30S subunit. The initiating AUG is guided to its precise position on the 30S subunit by an initiating signal called the Shine-Dalgarno sequence in the mRNA, centred 8–13 base pairs to the 5′ side of the initiating codon. Generally consisting of four-nine purine residues, the Shine-Dalgarno sequence is recognized by and base pairs with a complementary pyrimidine-rich sequence near the 3′-end of the 16S rRNA of the 30S subunit. This mRNA-rRNA interaction fixes the mRNA, so that the AUG is correctly positioned for initiation of translation.
 Ribosomes have two sites that bind aminoacyl-tRNAs, the aminoacyl or A-site and the Peptidyl or P-site. Both 30S and 50S subunits contribute to the characteristic of each site. The initiating AUG is positioned in the P-site, which is the only site to which fMet-tRNAfMet can bind. During subsequent elongation stage, all other aminoacyl-tRNAs bind to the A-site.
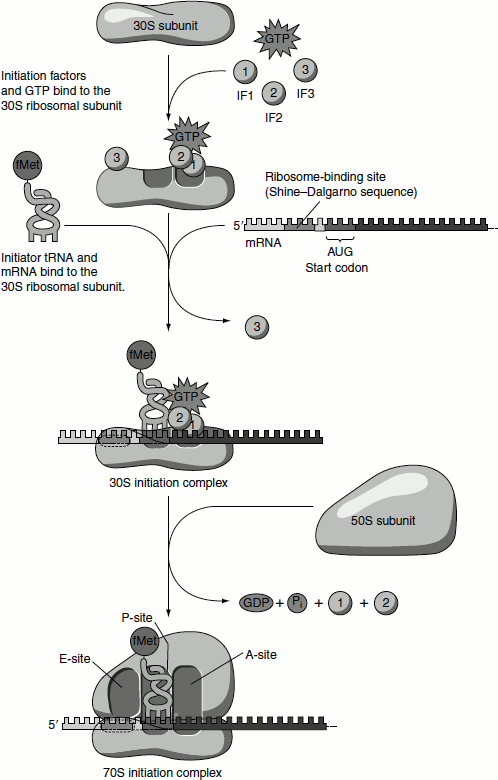
Ribosomes have two sites that bind aminoacyl-tRNAs, the aminoacyl or A-site and the Peptidyl or P-site. Both 30S and 50S subunits contribute to the characteristic of each site. The initiating AUG is positioned in the P-site, which is the only site to which fMet-tRNAfMet can bind. During subsequent elongation stage, all other aminoacyl-tRNAs bind to the A-site. - In the second step of the initiation process, the complex consisting of the 30S subunit, IF-3 and mRNA now forms a still larger complex by binding IF-2, which is already bound to GTP and the initiating fMet-tRNAfMet. The anticodon of this tRNA pairs correctly with the initiation codon.
- In the third step, this large complex combines with the 50S ribosomal subunit; simultaneously, the GTP molecule bound to IF-2 is hydrolysed to GDP and Pi (which are released). IF-3 and IF-2 also depart from the ribosome (Figure 5.9).
Elongation
Elongation involves the stepwise addition of amino acids to the polypeptide chain. Elongation requires:
- The initiation complex described above,
- The next aminoacyl-tRNA, specified by the next codon in the mRNA,
- A set of soluble cytosolic proteins called elongation factors (EFs) (EF-Tu, EF-Ts and EF-G)
- GTP.
Figure 5.9 Initiation of translation in prokaryotes
Elongation comprises three steps:
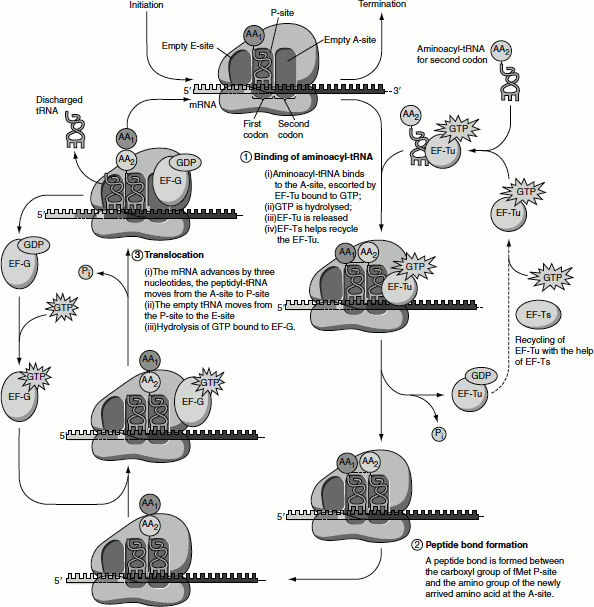
- In the first step of elongation, the next aminoacyl-tRNA is first bound to a complex of EF-Tu containing a molecule of GTP. The resulting aminoacyl-tRNA-EF-Tu-GTP complex is then bound to the A-site of the 70S initiation complex. The GTP is hydrolysed, an EF-Tu-GDP complex is released from the 70S ribosome and EF-Tu-GTP is regenerated. The guanine nucleotide exchange factor (GEF) EF-Ts mediates the regeneration of used form EF-Tu-GDP into the active form EF-Tu-GTP. First, EF-Ts displaces the GDP from EF-Tu, forming EF-Tu-EF-Ts. Then, GTP displaces EF-Ts reforming EF-Tu-GTP. This active complex binds aminoacyl-tRNA and the released EF-Ts can recycle. The hydrolysis of EF-Tu-GTP is relatively slow; it takes longer time for an incorrect aminoacyl-tRNA to dissociate from the A-site; therefore, ribosome checks the codon-anticodon interactions and most incorrect species are removed at this stage. Proof reading by the ribosome is made possible by the GTPase activity of Ef-Tu and thus makes an important contribution to the rate and fidelity of the overall biosynthetic process.
- In the second step, a new peptide bond is formed between the amino acids bound by their tRNAs to the A-site and P-site on the ribosome (Figure 5.10). This occurs by the transfer of the initiating formylmethionine group from its tRNA to the amino group of the amino acid now in the A-site. The α amino group of the amino acid in the A-site acts as a nucleophile, get attacks the carboxyl group of the amino acid in the P-site to form the peptide bond. This reaction produces a dipeptidyl-tRNAfMet in the A-site and now ‘uncharged’ (deacylated) tRNAfMet remains bound to the P-site. The activity responsible for the synthesis of peptide bond is called peptidyl transferase. Peptidyl transferase is a function of the large ribosomal subunit. The 23S rRNA has peptidyl transferase activity.
- The 3 step of elongation is called ‘translocation’ (Figure 5.11).The ribosome advances by three nucleotides along the mRNA towards the 3′-end. This movement of the ribosome shifts the dipeptidyl-tRNA from the A-site to P-site and the deacylated tRNA is released from the exit-site or E-site. The third codon of the mRNA will now be in the A-site and the second codon in the P-site. This shift of the ribosome along mRNA requires EF-G (also called translocase). The energy for the process is provided by the hydrolysis of GTP. The ribosome that switches between alternative and discrete co-formations results in the changes in rRNA base pairing, breaking many of its contact with the tRNA and allows the movement of the ribosome. EF-G is released following ribosome movement. Hydrolysis of GTP is needed to release EF-G.
Figure 5.10 Peptide bond formation
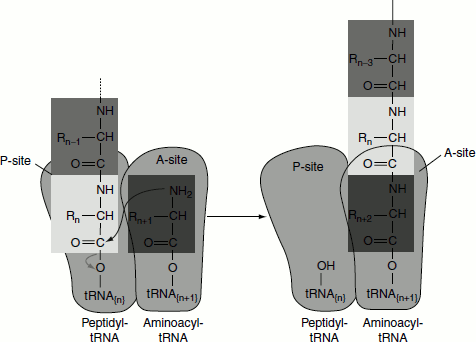
Figure 5.11 Elongation of the peptide chain
Termination
Elongation continues until the ribosome adds the last amino acid, completing the polypeptide coded by the mRNA. Termination is signalled by termination or stop codons in the mRNA (UAA, UAG and UGA), immediately following the last amino acid codon (Figure 5.12).
Figure 5.12 Termination of translation
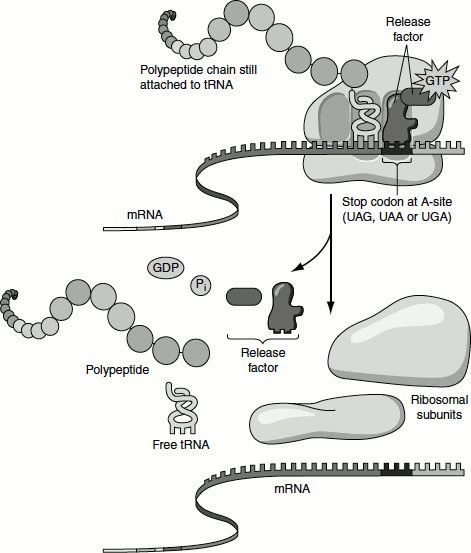
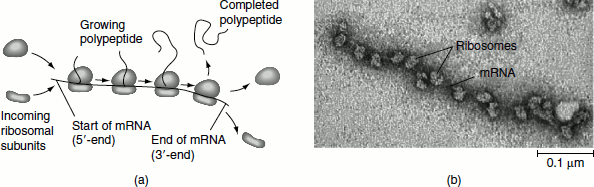
Figure 5.13 Polysomes (a) An mRNA molecule is generally translated simultaneously by several ribosomes in clusters called polyribosomes (b) This micrograph shows a large polyribo-some in a prokaryotic cell (TEM)
In bacterial genes, UAA is the most commonly used termination codon. UGA is used more frequently than UAG. The termination codons are recognized by release factors, namely RF1 and RF2 (class 1 release factors) and RF3 (class 2 release factors). RF1 recognizes UAA and UAG. RF2 recognizes UGA and UAA. The factors act at the ribosomal A-site and require peptidyl-tRNA at the P-site. The class 1 release factors are assisted by class 2 release factors, which are not codon-specific. The class 2 factors are GTP-binding proteins. In E. coli, the role of class 2 factor is to release class 1 factor from the ribosome. The class 1 factors recognize the termination codons and activate the ribosome to hydrolyse the peptidyl-tRNA. The peptidyl-tRNA transfers the growing peptide chain to a water molecule rather than to another amino acid. At this point, RF1 and RF2 are released by class 2 RF3 followed by the dissociation of the 70S ribosome into 50S and 30S ribosomes.
Polyribosomes are the Active Structures of Protein Synthesis
Active protein-synthesizing units consist of an mRNA with several ribosomes attached to it. Such structures are polyribosomes or simply called polysomes (Figure 5.13). All protein synthesis occurs on polysomes. In the polysome, each ribosome is traversing the mRNA and independently translating it into polypeptide. The farther a ribosome has moved along the mRNA, the greater the length of its associated polypeptide product. In prokaryotes, as many as 10 ribosomes may be found in a polysome. Ultimately, as many as 300 ribosomes may translate an mRNA, so as many as 300 enzyme molecules may be produced from a single transcript. Eukaryotic polysomes typically contain fewer than 10 ribosomes.