Besides being bound by two membranes, both mitochondria and chloroplasts also contain similar type of electrotransport proteins and use an F-class ATPase to synthesize ATP. The growth and division of mitochondria and chloroplasts are not coupled to nuclear division.
Proteins encoded by the mitochondrial and chloroplast DNA are synthesized on the ribosomes within the organelles and directed to the correct sub-compartment immediately after their synthesis. The majority of proteins in mitochondria and chloroplasts, however, are encoded by genes in the nucleus and are imported into the organelle after their synthesis in the cytosol. Proteins synthesized in the cytosol that are destined for the matrix of mitochondria or to the stroma of chloroplasts contain specific N-terminal uptake-targeting sequences that specify binding to receptor proteins on the organelle surface. This targeting sequence is cleaved once it reaches the matrix or stroma (Figure 6.11).
Transport of Proteins to Mitochondria
Proteins targeted from the cytosol to same mitochondrial destinations have targeting signal sequences that share common motif. Thus, the receptors that recognize these signals are able to bind a number of different but related sequences. One of the sequences for localizing proteins to the mitochondria is the ‘matrix-targeting sequences’. These sequences are located at the N-terminus and are about 20–25 amino acids in length. They are rich in hydrophobic amino acids, basic amino acids (arginine and lysine) and hydroxylated ones (serine and threonine). These sequences lack negatively charged amino acids (acidic amino acids) such as aspartate and glutamate.
Figure 6.11 Translocons of mitochondrion and chloroplast
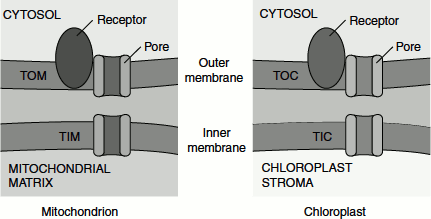
Mitochondrial ‘matrix-targeting sequences’ assume an a-helical conformation in which positively charged amino acids predominate on one side and hydrophobic amino acids predominate on the other side; thus, these sequences are amphipathic.
Transport of proteins to matrix of mitochondria
Proteins targeted to the mitochondria, soon after their synthesis interact directly with the mitochondrial membrane. Only unfolded proteins can be imported into the mitochondria. Chaperone proteins such as Hsc70 keep newly synthesized protein in an unfolded state. The proteins targeted to mitochondrial matrix takes a three-step route:
- First to the outer mitochondrial membrane,
- Second to the inner mitochondrial membrane and
- Finally to the mitochondrial matrix.
Transport to the outer mitochondrial membrane
The mitochondrial import of an unfolded protein is initiated by the binding of a mitochondrial sequence to an import receptor in the outer mitochondrial membrane. The N-terminal matrix-targeting sequences are recognized by proteins called Tom20 and Tom22. These are proteins in the outer mitochondrial membrane which are involved in targeting; Tom for translocon of the outer membrane.
The import receptor subsequently transfers the proteins to an import channel in the outer membrane. This channel is mainly composed of the Tom40 proteins. This is known as ‘import pore’, because all mitochondrial proteins gain access to the interior compartments of the mitochondria through this channel. Tom40 forms largely passive channel. These are wide enough to accommodate an unfolded protein, through the outer mitochondrial membrane.
Transport to the inner mitochondrial membrane
Transfer through the outer membrane occurs simultaneously with the transfer through an inner membrane channel. The inner membrane channel is composed of proteins called Tim23 and Tim17 proteins (Tim for translocon of inner membrane) (Figure 6.12).
Figure 6.12 Protein targeting to mitochondrial inter membrane space
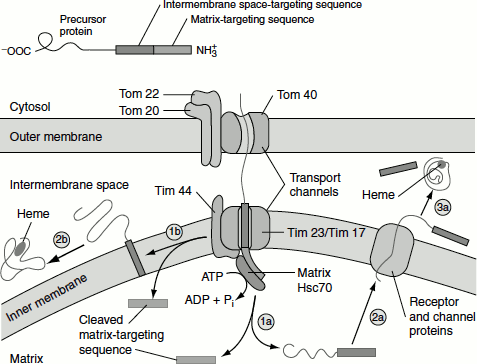
(1a) Protease cleavage of matrix targeting sequence.
(1b) Protease cleavage of matrix targeting sequence.
(2a) Transport of cleaved protein with intermembrane space targeting sequence through channel proteins.
(2b) Transmembrane insertion of the intermembrane space targeting sequence; cleavage and release of the protein into the intermembrane space.
(3a) Cleavage of the intermembrane space targeting sequence and release into intermembrane space.
Transport to the mitochondrial matrix
Translocation into the mitochondrial matrix occurs at ‘contact site’ where outer and inner membranes are in close proximity. Soon after the AMterminal matrix-targeting sequence enters the mitochondrial matrix, it is removed by the protease that is present within the matrix. The emerging protein is also bound by the protein that is present in the translocation channels in the inner mitochondrial membrane known as matrix Hsc70. This binding requires interaction with Tim44. This interaction stimulates ATP hydrolysis by matrix Hsc70 which powers the translocation of proteins into the matrix. Some imported proteins can fold into their final active conformation without further assistance. However, many proteins require the help of a chaperonin for their final folding (Figure 6.13).
Transport of proteins to inner mitochondrial membrane
Three separate pathways are known to target proteins to the inner mitochondrial membrane.
One pathway uses the same machinery that is used for targeting of matrix proteins. A cytochrome oxidase subunit called Cox Va is a typical protein transported by this pathway.
Figure 6.13 Protein targeting to mitochondrial matrix
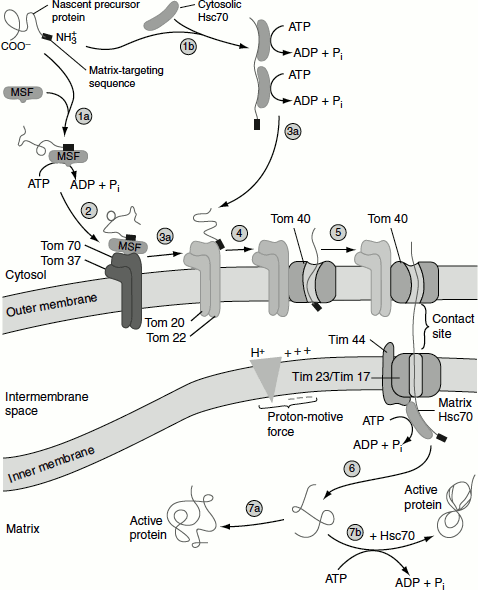
(1a, 2) Matrix targeting sequence directing the protein to outer membrane.
(1b) Binding of the protein with chaperonin Hsc70.
(3a, 4, 5) Transport of the protein through translocation channel powered by the energy of hydrolysis of ATP.
(6, 7a) Release of the protein into the matrix.
(7b) Hsc70 assisted folding of the protein
The second pathway to the inner membrane involves the use of matrix-targeting sequence and an internal hydrophobic domains that are recognized by an inner membrane protein termed Oxa1. The proteins are translocated (at least a portion of the protein) into the matrix through the Tom20/Tom22 and Tim23/17 channels.
After the cleavage of the matrix-targeting sequence, the protein is inserted into the inner membrane. This process requires the interaction with Oxa1 and other inner membrane proteins. Oxa1 also participates in the inner membrane insertion of certain proteins (e.g., subunit II of cytochrome oxidase) that are encoded by mitochondrial DNA and synthesized in the matrix by mitochondrial ribosome.
The final pathway for insertion in the inner mitochondrial membrane is followed by multipass proteins that contain six membrane-spanning domains such as ADP/ATP antiporter. These proteins, which lack the usual N-terminal matrix-targeting sequence, contain multiple internal mitochondrial targeting sequence. After the internal sequences are recognized by Tom70, the imported protein pass through the outer membrane through the general import pore. The protein is then transferred to a second translocation complex in the inner membrane that is composed of Tim9 and Tim10, which reside in the inner membrane space. These act as chaperones to guide imported protein from the general import pore to the Tim22/54 complex which incorporates the multiple hydrophobic segments of the imported protein into the inner membrane.
Transport of proteins to inter membrane space
The space that is present between the outer and inner mitochondrial membranes is called the intermembrane space. Two pathways deliver cytosolic proteins to the intermembrane space.
One pathway involves the use of an N-terminal sequence (hydrophobic sequence) and an N-terminal matrix-targeting sequence. Both of these sequences are finally cleaved. The N-terminal sequence (hydrophobic sequence) blocks the complete translocation of the protein across the inner membrane. The hydrophobic sequence after embedding the protein then diffuses laterally away from Tim23/17 translocation channel.
A protease in the inner membrane cleaves the protein near the hydrophobic transmembrane segment thus releasing the mature protein in the soluble form into the intermembrane space.
Cytochrome C heme lyase, the enzyme responsible for the attachment of heme to cytochrome C, explains the second pathway for targeting to the intermembrane space. In this pathway, imported protein is delivered directly to the intermembrane space via the general import pore without the involvement of any inner membrane translocation factors. Since translocation through the Tom40 general import pore is not coupled to any energetically favourable process such as the hydrolysis of ATP or GTP, the unidirectional translocation through the outer membrane is unclear. One possibility is that cytochrome C heme lyase passively diffuse through the outer membrane and then is trapped within the intermembrane space by binding to another protein that is delivered to that location by one of the translocation mechanisms discussed previously.
Transport of proteins to outer mitochondrial membrane
A short matrix-targeting sequence at the N-terminus of mitochondrial porin (P70) followed by a long stretch of hydrophobic sequence is involved in this translocation. Long stretch of hydrophobic sequence functions as a stop-transfer sequence that prevents the transfer of the protein into the matrix and anchors it as an integral protein in the outer membrane.
Transport of Proteins to Chloroplast
The chloroplasts have three membranes: outer membrane, inner membrane and the thylakoid membrane; consequently, the chloroplasts have three membrane spaces, which are intermembranous space, the stroma and the thylakoid membrane space. Proteins need to go through each of these membranes and into their respective compartments.
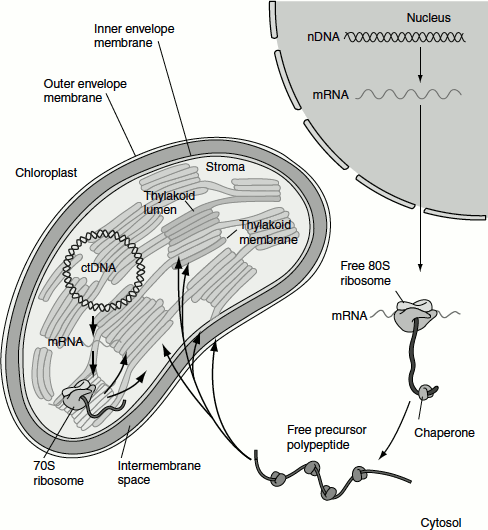
Figure 6.14 Protein targeting in plant cell (a) Free ribosomes in cytosol (b) Membrane-bound ribosomes
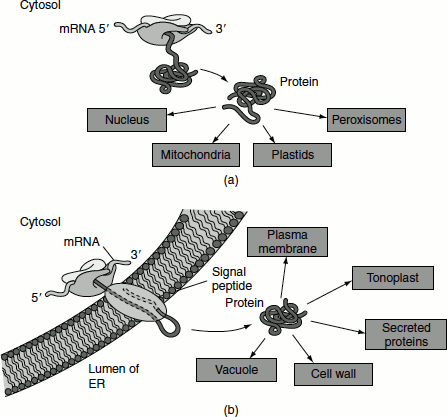
Proteins synthesized on membrane-bound ribosomes are first translocated into the lumen of the ER and then transported to the Golgi apparatus. These proteins may subsequently be targeted to the plasma membrane or the tonoplast, secreted or sent to the vacuole (Figure 6.14).
Chloroplast proteins may be encoded by nuclear DNA or chloroplast DNA; the respective mRNAs are translated by ribosomes in the cytosol (80S ribosome) or in the chloroplast stroma (70S ribosome). Proteins made as a precursor polypeptide in the cytosol may be targeted to the outer membrane or may enter the chloroplast stroma, to the thylakoid membrane, thylakoid lumen or inner envelope membrane (Figure 6.15).
Mechanism of protein import into the chloroplast
The import of proteins from the cytosol to chloroplasts shares several characteristics with mitochondrial import. In both processes, the imported proteins are synthesized as cytosolic precursors containing N-terminal uptake-targeting sequences that direct each protein to its correct subcompartment and are subsequently cleaved. Protein import from the cytosol into the chloroplast stroma (equivalent to the mitochondrial matrix) occurs, as in mitochondria, at the points where the outer and inner organelle membranes are in close contact. Finally, the protein import into both organelles requires energy. Despite the similarities just noted, the mechanisms of chloroplast and mitochondrial protein import differ in various ways.
Figure 6.15 Protein targeting to chloroplast sub compartments
Protein transport to chloroplast stromal space
Among the proteins found in the chloroplast stroma are the enzymes of the Calvin cycle. The large (L) subunit of ribulose-1,5-bisphosphate carboxylase (RuBisCO) is encoded by chloroplast DNA and synthesized on chloroplast ribosomes in the stromal space. The small (S) subunit of RuBisCO and all the other Calvin-cycle enzymes are encoded by nuclear genes and transported to chloroplasts after their synthesis in the cytosol (Figure 6.16).
The S-subunit of RuBisCO is synthesized on free cytosolic polyribosomes in a precursor form that has an N-terminal stromal-import sequence of about 44 amino acids. It is maintained in an unfolded state by binding to cytosolic chaperones; they can import the S-subunit precursor after its synthesis. After the unfolded precursor enters the stromal space, it binds transiently to a stromal Hsc70 chaperone, and the N-terminal sequence is cleaved. In the reactions that are facilitated by Hsc60 chaperonins, eight S-subunits combine with the eight L-subunits to yield the active RuBisCO enzyme.
Figure 6.16 Protein targeting to stroma of chloroplast
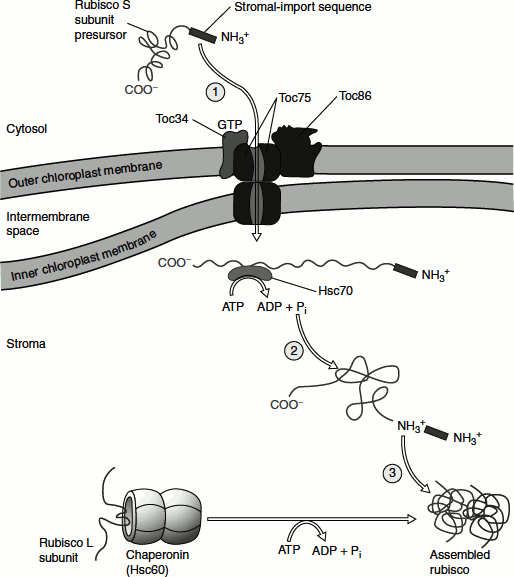
The import process involves the use of three chloroplast outer-membrane proteins, namely:
- A receptor that binds the stromal-targeting sequence,
- A transport channel protein and
- Five inner-membrane proteins.
Proteins are imported in the unfolded state into the stroma. The import process requires ATP hydrolysis that is catalysed by stromal chaperones, which functions similarly to Hsc70 of the mitochondrial matrix. Unlike mitochondria, however, chloroplasts cannot generate an electrochemical gradient (proton-motive force) across their inner membrane. Thus, protein import into the chloroplast stroma appears to be powered solely by ATP hydrolysis.
Targeting to the thylakoids
The targeting of proteins to thylakoid membrane or to the lumen involves the following steps (Figure 6.17):
- Transport to both the outer and inner chloroplast membranes to enter the stroma,
- Transport through the stroma and
- Transported protein can either be inserted into the thylakoid membrane or cross that membrane and enter the thylakoid lumen.
Figure 6.17 Protein targeting to thylakoids
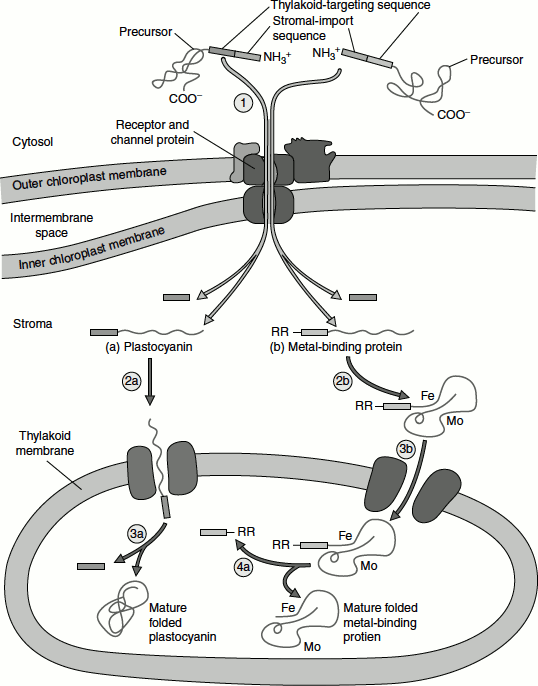
Proteins that are destined for the thylakoid lumen, such as plastocyanin, require the successive action of two targeting sequences:
- The first sequence targets the protein to the stroma.
- The second sequence targets the protein from the stroma to the thylakoid lumen.
Four separate thylakoid-import systems, each transporting a different set of proteins from the stroma into the thylakoid lumen, have been identified.
One of the import systems is similar to the ER import and this system functions even in the absence of a pH gradient across the thylakoid membrane.
The other system involves the use of the thylakoid-membrane protein Hef106 which assists in translocating folded proteins and their bound cofactors into the thylakoid lumen. The protein uptake is powered by the pH gradient normally maintained across the thylakoid membrane.