At the Level of Post-transcriptional Modification
In eukaryotes, much of the gene regulation takes place during the mRNA processing. Approximately 75 per cent of pre-mRNAs are degraded within the nucleus. This selective degradation acts as the control point of gene regulation. Alternative splicing of pre-mRNA permits the synthesis of different versions of a protein. Alternative splicing also affect the outcome of the development processes.
By Controlling the Stability of mRNA Molecule
All mRNA molecules have a characteristic life span. They are degraded in the cytoplasm. Some are degraded within minutes after their synthesis some last hours or even months or years, as in the case of the mRNAs stored in oocytes. The poly(A) tail and the 5′ guanosine caps protect the mRNAs from the action of nucleases and thereby increase their life span. Histone mRNAs that lack a poly(A) tail, a stem and loop structure is formed at the 3′ end.
At Translational Level
Some genes can be regulated at translational level. For example, in some anucleate cells, such as reticulocytes, transcriptional control is unavailable and hence regulated at translational level.
The immunoglobulin, IgM, exists in two forms namely a membrane-bound form and secreted form. Both the forms of the protein are encoded by the same gene and differ only in their heavy chains carboxyl ends. The IgM gene has two poly(A) addition sites and has a number of exons that can produce two alternative forms. The transmembrane anchor of the membrane-bound form is encoded by exons 5 and 6. The secreted product is produced when the exon 4 splice is not made and these transcripts are terminated just after exon 4.
At Post-translational Level
In some cases, regulation occurs after translation. The tubulin mRNA gene expression is regulated post-translationally. The first four amino acids, i.e., met, arg, glu and ile, of the tubulin gene product constitute a recognition element to which regulatory factors bind and degrade the mRNA depending on tubulin concentration in the cytoplasm.
By Hormone Action
Just as in prokaryotes, eukaryotes have many systems for responding to specific molecules with which they come in contact. For example, the hormone oestrogen, which is released by the ovaries, is cholesterol-derived steroid hormones. They are required for the development of female secondary sexual characteristics. As oestrogens are hydrophobic, they can cross the cell membrane. Inside the cell, they bind to soluble receptor proteins. Oestrogen receptor proteins are members of the large family of proteins that act as receptors for a wide range of hydrophobic molecules, including other steroid hormones, thyroid hormones and retinoids.
On binding the signal molecule (ligand), the ligand–receptor complex modifies the expression of specific genes by binding to control elements in the DNA called ‘hormone response elements’. The hormone-binding receptors usually have two domains; the DNA-binding domain and the ligand-binding domain. The DNA-binding domain has a zinc-finger motif.
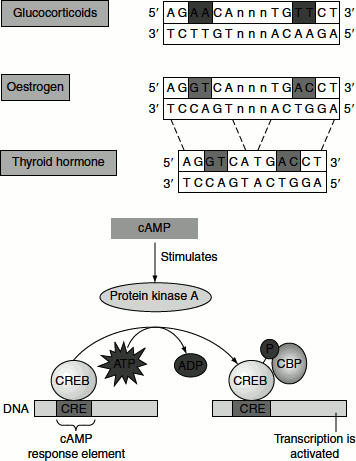
Oestrogen receptors bind to specific DNA sites referred to as oestrogen response elements or EREs that contain the consensus sequence 5′AGGTCANNNTGACCT3′. Oestrogen receptor binds to such sites as a dimer.
The binding of the ligand receptor complex to the hormone response elements results in gene regulation, i.e., either activates or inactivates gene expression depending on the fact whether it is complexed with activator or repressor protein. For example, oestrogen binding to the receptor stimulates co-activator binding and up regulates gene expression.
Receptors for thyroid hormone and retinoic acid repress the transcription in the absence of the ligand. In their unbound form, the ligand-binding domains of these receptors bind to co-repressors. Examples of such co-repressor proteins include silencing mediator for retinoid and thyroid hormone receptors (SMRT) and nuclear hormone co-repressor (N-cor). Ligand binding triggers the release of co-repressor and frees the ligand-binding domain for binding to a co-activator.
Ligand binding to membrane receptors can regulate transcription through phosphorylation cascades. For example, the binding of epinephrine to 7TM receptors results in the activation of a G protein. Activated G protein binds to and activates adenylate cyclase, increasing the concentration of cAMP. cAMP binds to the regulatory subunits of protein kinase A (PKA) activating the enzyme. PKA also phosphorylates the cAMP response element-binding protein (CREB), which is a transcription factor, that binds specific DNA sequences as a dimer. Phosphorylated CREB binds to co-activator protein termed cAMP-binding protein (CBP).
Thus, the pathway initiated by epinephrine binding includes the phosphorylation of a transcription factor, the recruitment of co-activator and the assembly of complexes that participate in chromatin remodelling and transcription initiation.