There are four types of RNA, each encoded by its own type of gene:
- mRNA—Messenger RNA: Encodes amino acid sequence of a polypeptide.
- tRNA—Transfer RNA: Brings amino acids to ribosomes during the translation.
- rRNA—Ribosomal RNA: With ribosomal proteins makes up the ribosomes, the organelles that translate the mRNA.
- snRNA—Small nuclear RNA: With proteins forms complexes that are used in RNA processing in eukaryotes (not found in prokaryotes).
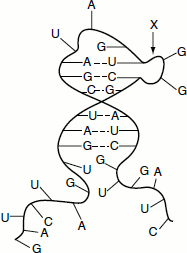
Figure 1.9 Structure of a eukayotic mRNA

Messenger RNA
‘The genetic information stored in the DNA, inside the nucleus, is conveyed to the ribosomes in the cytosol for the synthesis of proteins through the mRNA. Thus, the mRNA carries the genetic message from the nucleus to the cytosol.’ The genetic code is translated for mRNA, not for DNA. The structure of mRNA includes more than simply a copy of the gene from the DNA. On one end of the mRNA is a cap. This is a structure that allows the mRNA to bind to the ribosome and is very important in the protein synthesis. mRNA contains a section of RNA that is non-coding. This section can vary in length. Next is an initiation codon, which signals the beginning of the coding sequence, and ends with the stop codon. Finally, there is the coding region that contains the copy of the genes. The mRNA also bears a series of adenine residue at its 3′ end called the poly-A tail (Figure 1.9).
Ribosomal RNA
Ribosomal RNA (rRNA) or insoluble RNA constitutes the largest part (up to 80%) of the total cellular RNA. It is found primarily in the ribosomes although; it is synthesized in the nucleus. It is also detected in the nucleus. It contains the four major RNA bases with a slight degree of methylation and shows differences in the relative proportions of the bases between species. rRNA molecules are single polynucleotide strands that are unbranched and flexible. At low ionic strength, rRNA behaves as a random coil; however, with increasing ionic strength, the molecule takes secondary structures showing helical regions produced by base pairing between self-complementary sequences, adenine and uracil, guanine and cytosine (Figure 1.10).
Figure 1.10 Structure rRNA
Types of rRNA
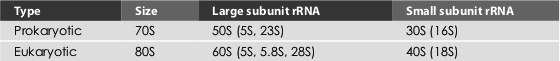
The eukaryotic cells have three kinds of rRNA molecules, namely 28S rRNA (the sedimentation constant varies between 25S and 30S depending up on the species), 18S and 5S rRNAs. The 28S and 5S rRNAs occur in 60S ribosomal submit, while 18S rRNA occurs in 40S ribosomal submit of the 80S ribosomes of eukaryotes. The prokaryotes contain 5S and 23S rRNAs in the 50S subunit and 16S in the 30S subunit (Table 1.5).
The sequences of the small and large rRNAs from several thousand organisms are now known. Though their primary nucleotide sequences vary considerably, the same parts of each type of rRNA theoretically can form base-paired stem-loops, generating a similar three-dimensional structure for each rRNA in all organisms.
Table 1.5 Prokaryotic and eukaryotic rRNAs
Transfer RNA
These are small RNA molecules and consist of about 73 to 94 nucleotides in a single chain. They are also known as soluble RNAs and help in transferring amino acids from the ‘amino acid pool’ to the site of protein synthesis. tRNAs account for 15 per cent of the total RNA of the cell. Eukaryotic cells are estimated to have approximately 60 different types of tRNAs. tRNA acts as an ‘adapter molecule’ linking the information in mRNA codons with the specific amino acids in proteins. For each of the 20 amino acids, there is at least one specific type of tRNA molecule.
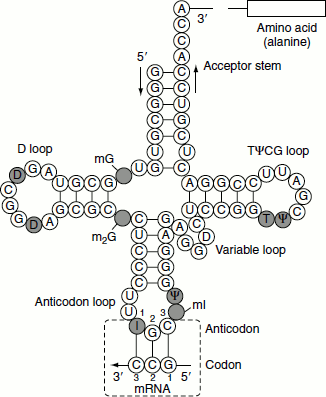
Robert Holley et al. worked out the complete sequence of yeast alanine tRNA molecule. All tRNA molecules have a two-dimensional cloverleaf structure. This is because the majority of the bases are hydrogen-bonded to one another; the complementary stretches of bases in the chain form stem and loop structures and hence the overall pattern of H-bonding can be represented as a cloverleaf. Each cloverleaf consists of four H-bonded segments—three loops and the stem where the 3′- and 5′-ends of the molecule meet. These four segments are designated ‘the acceptor stem, the dihydrouridine loop, the anticodon loop and the ribothymidine pseudouracil cytosine loop’. The four stems are short double helices stabilized by Watson–Crick base pairing; three of the four stems have loops containing seven or eight bases at their ends, while the unlooped stem contains the free 3′ and 5′ ends of the chain (Figure 1.11).
Figure 1.11 tRNA cloverleaf structure
The acceptor stem
Specific aminoacyl-tRNA synthetases recognize the surface structure of each tRNA for a specific amino acid and covalently attach the proper amino acid to the unlooped ‘amino acid acceptor stem’. The 3′ end of all tRNAs has the sequence CCA, which is added after the synthesis and processing of the tRNA are completed. The 3′-OH of the adenine residue of the CCA terminus is esterified with the carboxyl group of the amino acid.
The anticodon loop
Three nucleotides termed the anticodon are located at the centre of the ‘anticodon loop’. These can form base pairs with the three complementary nucleotides forming a codon in mRNA. Viewed in three dimensions, the folded tRNA molecule has an L shape with the anticodon loop and acceptor stem forming the ends of the two arms. Base pairing of anticodon with the codon on mRNA allows a particular tRNA species to deliver its amino acid to the protein-synthesizing apparatus. It represents the key event in translating the information in the nucleic acid sequence, so that the appropriate amino acid is inserted at the right place in the amino acid sequence of the protein being synthesized.
The dihydrouridine loop
Ribosomes bind tRNAs through the recognition of this loop. The ‘D loop’ is so named because this tRNA loop often contains dihydrouridine (D) residues. In addition to dihydrouridine, tRNAs characteristically contain a number of unusual bases, including inosine, thiouridine, pseudouridine and hypermethylated purines.
The ribothymidine pseudouracil cytosine loop
Most tRNAs are synthesized with a four-base sequence of UUCG near the middle of the molecule. The first uridylate is methylated to become a thymidylate; the second is rearranged into a pseudouridylate (abbreviated ψ), in which the ribose is attached to carbon 5 instead of to nitrogen 1 of the uracil. These modifications produce a characteristic ‘TψCG loop’.
Extra arm or variable loop
The next loop in tRNA sequence in the 5′ → 3′ direction is a loop that varies from tRNA to tRNA in the number of residues that it has and is called ‘extra arm or variable loop’. Some tRNAs lack this loop.
tRNA tertiary structure
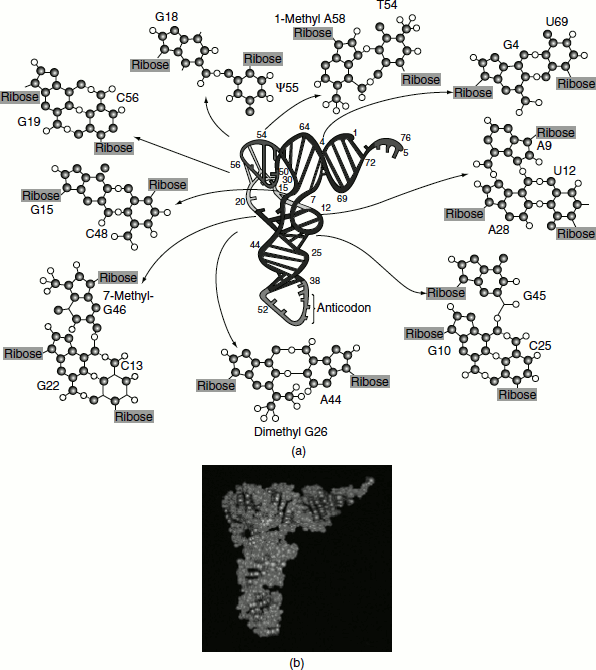
Tertiary structure in tRNA arises from hydrogen-bonding interactions between bases in the D loop with bases in the variable and ‘TψCG loops’, as shown for yeast phenylalanine tRNA (Figure 1.12).
These H bonds fold the D loop and ‘TψCG loop’ together and bend the cloverleaf into the stable L-shaped tertiary form. Many of these H bonds involve base pairs that are not canonical A=T or G≡C pairings. The amino acid acceptor stem is at one end of the L, separated by approximately 7 nm from the anticodon at the opposite end of the L. The D loop and ‘TψCG loop’ form the corner of the L. In the L-conformation, the bases are oriented to maximize hydrophobic stacking interactions between their flat faces. Such stacking interactions stabilize L-form tertiary structure of the tRNA.
Figure 1.12 (a) The three–dimensional structure of yeast phenylalanine tRNA as deducted from X-ray diffraction studies of its crystals. The tertiary folding is illustrated in the center of the diagram with the ribose—phosphate backbone presented as a continuous ribbon; H bonds are indicated by crossbars. Unpaired bases are shown as short, unconnected rods. The anticodon loop is at the bottom and the —CCA 3′ —OH acceptor end is at the top right. The various types of noncanonical hydrogen-bonding interactions observed between bases surround the central molecule. Three of these structures show examples of unusual H-bonded interactions involving three bases; these interactions aid in establishing tRNA tertiary structure. (b) A space-filling model of the molecule. (After Kim, S. H., in Schimmel, P., Söll, D., and Abelson, J. N., eds., 1979. Transfer RNA: Structure, Properties, and Recognition. New York: Cold Spring Harbor Laboratory)